-
PDF
- Split View
-
Views
-
Cite
Cite
Qingchen Rui, Xiaoyun Tan, Feng Liu, Yanbin Li, Xin Liu, Bingxuan Li, Junxia Wang, Huiting Yang, Lixin Qiao, Tingting Li, Shuo Fang, Rongrong Gao, Wang Wang, Sebastian Y Bednarek, Yiqun Bao, Syntaxin of plants31 (SYP31) and SYP32 is essential for Golgi morphology maintenance and pollen development, Plant Physiology, Volume 186, Issue 1, May 2021, Pages 330–343, https://doi.org/10.1093/plphys/kiab049
Close - Share Icon Share
Abstract
Pollen development is a key process for the sexual reproduction of angiosperms. The Golgi plays a critical role in pollen development via the synthesis and transport of cell wall materials. However, little is known about the molecular mechanisms underlying the maintenance of Golgi integrity in plants. In Arabidopsis thaliana, syntaxin of plants (SYP) 3 family proteins SYP31 and SYP32 are the only two Golgi-localized Qa-soluble N-ethylmaleimide sensitive factor attachment protein receptors (SNAREs) with unknown endogenous functions. Here, we demonstrate the roles of SYP31 and SYP32 in modulating Golgi morphology and pollen development. Two independent lines of syp31/+ syp32/+ double mutants were male gametophytic lethal; the zero transmission rate of syp31 syp32 mutations was restored to largely normal levels by pSYP32:SYP32 but not pSYP32:SYP31 transgenes, indicating their functional differences in pollen development. The initial arrest of syp31 syp32 pollen occurred during the transition from the microspore to the bicellular stage, where cell plate formation in pollen mitosis I (PMI) and deposition of intine were abnormal. In syp31 syp32 pollen, the number and length of Golgi cisterna were significantly reduced, accompanied by many surrounding vesicles, which could be largely attributed to defects in anterograde and retrograde trafficking routes. SYP31 and SYP32 directly interacted with COG3, a subunit of the conserved oligomeric Golgi (COG) complex and were responsible for its Golgi localization, providing an underlying mechanism for SYP31/32 function in intra-Golgi trafficking. We propose that SYP31 and SYP32 play partially redundant roles in pollen development by modulating protein trafficking and Golgi structure.
Introduction
Male gametophyte development is a key process for plant reproduction. The development of Arabidopsis thaliana male gametophytes starts from haploid microspores formed by meiosis of microsporocytes. After being released from the tetrads by degrading the surrounding callose walls (Owen and Makaroff, 1995), each microspore undergoes an asymmetric pollen mitosis I (PMI) division to produce bicellular pollen (BCP) composed of a large vegetative cell and a small generative cell (Yamamoto et al., 2003). The tricellular pollen (TCP) with one vegetative and two sperm cells forms after another symmetric pollen mitosis II (PMII) division and eventually develop into mature pollen (MP).
Soluble N-ethylmaleimide sensitive factor attachment protein receptors (SNARE) proteins are well-defined machinery for the fusion between approaching vesicles and their target membrane. Based on the conserved residues located at the center of the SNARE motif, SNAREs are grouped into R-SNAREs and Q-SNAREs, and Q-SNAREs are further subdivided into four subfamilies: Qa-, Qb-, Qc-, and Qbc-SNAREs (Hong, 2005). Membrane fusion requires four SNARE motifs of cognate SNARE proteins that are anchored to the donor or acceptor membrane to form a four-helix bundle, which powers the membrane fusion (Jahn and Scheller, 2006).
Until now, only a handful of vesicle trafficking components have been implicated in pollen grain development. Pollen abortion occurs in the sec24b sec24c double mutant, where two coat protein complex II (COPII) structure proteins are disrupted (Tanaka et al., 2013). Loss of SEC22, an R-SNARE located at endoplasmic reticulum (ER), causes the plasma membrane-targeted Qa-SNARE SYP124 to be retained in the ER and impairs pollen development (El-Kasmi et al., 2011). In addition, several components of clathrin-mediated endocytosis (CME) are also required for male gametogenesis, such as clathrin light chain1 (CLC1; Wang et al., 2013), several subunits of the TPLATE complex (Van Damme et al., 2006; Gadeyne et al., 2014), and Dynamin-related proteins DRP1C and DRP2A/B (Kang et al., 2003; Backues et al., 2010). Interestingly, drp1c-1 mutant pollen displayed irregular plasma membrane and intine morphology, and collapsed during desiccation (Kang et al., 2003). drp2ab double mutant pollen exhibited branched or irregular cell plates during PMI (Backues et al., 2010). Recently, overexpressed Arabidopsis Tomosyn (AtTMS), a negative regulator of a set of plasma membrane-localized Qa-SNAREs, was shown to arrest pollen development where the intine deposition and cell plate formation during PMI were disrupted (Li et al., 2019). Apparently, more components of the vesicle trafficking machinery required for pollen development remain to be studied.
In plant cells, the Golgi apparatus is central to the secretory pathway and produces cell wall polysaccharides. Anterograde trafficking from ER to Golgi mediated by COPII vesicles is counterbalanced by the retrograde trafficking of COPI vesicles (Stefano et al., 2006; Paul and Frigerio, 2007). The conserved oligomeric Golgi (COG) complex is a Golgi-localized tethering complex; mutations in COG3 and COG8 subunits impair the COPI trafficking and Golgi integrity in pollen (Tan et al., 2016). However, little is known about how COG complex-dependent vesicle trafficking is operated.Syntaxin of plants 31 (SYP31) and SYP32 are the only two Golgi localized Qa-SNAREs in Arabidopsis (Sanderfoot et al., 2000; Uemura et al., 2004). Overexpression of SYP31 inhibits anterograde trafficking of a number of secretory markers in Nicotiana benthamiana protoplasts (Bubeck et al., 2008). SYP31 is proposed to play a role in Golgi reformation after Brefeldin A (BFA) removal (Ito et al., 2012; Ito et al., 2018). In addition, SYP31 localizes to the forming cell plate in Arabidopsis suspension cells, suggesting its involvement in the formation of the cell plate during somatic cytokinesis (Rancour et al., 2002). Based on these evidences, the role of SYP31 in ER–Golgi trafficking is speculated. Until now, there is no data indicating SYP31 in intra-Golgi transport. Moreover, the endogenous physiological functions of SYP31 and SYP32 in Arabidopsis remain unknown.
In this study, using genetic, cytological, and biochemical assays, we show that SYP31 and SYP32 play partially redundant roles in pollen development. Here, were explored detailed phenotype analyses of syp31/+ syp32/+ double mutants and the underlying molecular mechanisms.
Results
syp31 syp32 double mutations lead to male sterility
The SYP3 family in Arabidopsis consists of SYP31 and SYP32, which share 44% sequence identity at the amino acid level. We set out to characterize the T-DNA insertion lines of these two genes, syp31/+, syp32-1/+, and syp32-2/+ (Figure 1A). The homozygous syp31/− knockout mutant identified by PCR-based genotyping exhibited no noticeable phenotype (Supplemental Figure S1), and no male or female transmission defects were detected upon reciprocal crosses of the syp31/+ mutant (Table 1).
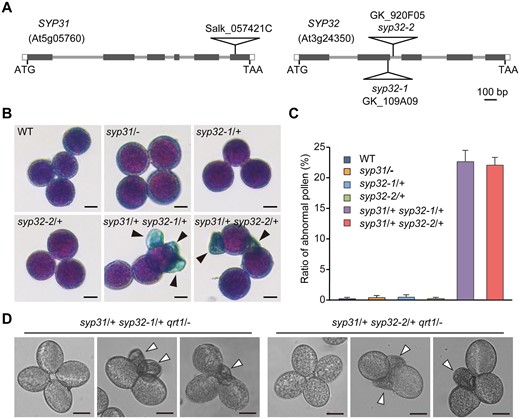
The double mutations in SYP31 and SYP32 cause pollen abortion. A, Schematic diagram of the SYP31 and SYP32 gene structures and T-DNA insertion sites. Black boxes represent exons. Gray lines represent introns. White boxes represent untranslated regions (UTRs). Triangles indicate T-DNA insertion sites. B, Alexander staining of MP. Arrows indicate aborted pollen. C, Statistical analysis of abnormal pollen from WT and indicated mutants (n > 500 pollen from three individual plants was counted for each line), values represent the means SD. D, Quartets from the syp31/+ syp32/+ qrt1/− triple mutant lines. Arrowheads indicate deformed pollen. Bars = 10 μm in (B) and (D).
| Crosses (Female × Male) . | Genotype of progeny . | TEF . | TEM . | |||||
|---|---|---|---|---|---|---|---|---|
| WT . | 31/+ . | 32-1/+ . | 32-2/+ . | 31/+32-1/+ . | 31/+32-2/+ . | |||
| 31/+ × +/+ | 48 | 48 | 100% | NA | ||||
| +/+ × 31/+ | 49 | 47 | NA | 96% | ||||
| 32-1/+ × +/+ | 478 | 6 | 1.3% | NA | ||||
| +/+ × 32-1/+ | 313 | 175 | NA | 56% | ||||
| 32-2/+ × +/+ | 152 | 0 | 0 | NA | ||||
| +/+ × 32-2/+ | 114 | 78 | NA | 68% | ||||
| +/+ × 31/+ 32-1/+ | 96 | 87 | 50 | 0 | NA | 0 | ||
| +/+ × 31/+ 32-2/+ | 89 | 80 | 32 | 0 | NA | 0 | ||
| Crosses (Female × Male) . | Genotype of progeny . | TEF . | TEM . | |||||
|---|---|---|---|---|---|---|---|---|
| WT . | 31/+ . | 32-1/+ . | 32-2/+ . | 31/+32-1/+ . | 31/+32-2/+ . | |||
| 31/+ × +/+ | 48 | 48 | 100% | NA | ||||
| +/+ × 31/+ | 49 | 47 | NA | 96% | ||||
| 32-1/+ × +/+ | 478 | 6 | 1.3% | NA | ||||
| +/+ × 32-1/+ | 313 | 175 | NA | 56% | ||||
| 32-2/+ × +/+ | 152 | 0 | 0 | NA | ||||
| +/+ × 32-2/+ | 114 | 78 | NA | 68% | ||||
| +/+ × 31/+ 32-1/+ | 96 | 87 | 50 | 0 | NA | 0 | ||
| +/+ × 31/+ 32-2/+ | 89 | 80 | 32 | 0 | NA | 0 | ||
For single mutants, TE = (progeny with insertion/progeny without insertion) ×100%; for double mutants: TE = (progeny with both insertions/progeny without insertion) ×100%; TEF and TEM, female and male transmission efficiency, respectively.
NA, not applicable; TE, transmission efficiency.
| Crosses (Female × Male) . | Genotype of progeny . | TEF . | TEM . | |||||
|---|---|---|---|---|---|---|---|---|
| WT . | 31/+ . | 32-1/+ . | 32-2/+ . | 31/+32-1/+ . | 31/+32-2/+ . | |||
| 31/+ × +/+ | 48 | 48 | 100% | NA | ||||
| +/+ × 31/+ | 49 | 47 | NA | 96% | ||||
| 32-1/+ × +/+ | 478 | 6 | 1.3% | NA | ||||
| +/+ × 32-1/+ | 313 | 175 | NA | 56% | ||||
| 32-2/+ × +/+ | 152 | 0 | 0 | NA | ||||
| +/+ × 32-2/+ | 114 | 78 | NA | 68% | ||||
| +/+ × 31/+ 32-1/+ | 96 | 87 | 50 | 0 | NA | 0 | ||
| +/+ × 31/+ 32-2/+ | 89 | 80 | 32 | 0 | NA | 0 | ||
| Crosses (Female × Male) . | Genotype of progeny . | TEF . | TEM . | |||||
|---|---|---|---|---|---|---|---|---|
| WT . | 31/+ . | 32-1/+ . | 32-2/+ . | 31/+32-1/+ . | 31/+32-2/+ . | |||
| 31/+ × +/+ | 48 | 48 | 100% | NA | ||||
| +/+ × 31/+ | 49 | 47 | NA | 96% | ||||
| 32-1/+ × +/+ | 478 | 6 | 1.3% | NA | ||||
| +/+ × 32-1/+ | 313 | 175 | NA | 56% | ||||
| 32-2/+ × +/+ | 152 | 0 | 0 | NA | ||||
| +/+ × 32-2/+ | 114 | 78 | NA | 68% | ||||
| +/+ × 31/+ 32-1/+ | 96 | 87 | 50 | 0 | NA | 0 | ||
| +/+ × 31/+ 32-2/+ | 89 | 80 | 32 | 0 | NA | 0 | ||
For single mutants, TE = (progeny with insertion/progeny without insertion) ×100%; for double mutants: TE = (progeny with both insertions/progeny without insertion) ×100%; TEF and TEM, female and male transmission efficiency, respectively.
NA, not applicable; TE, transmission efficiency.
In contrast, the progenies from self-pollinated syp32-1/+ and syp32-2/+ mutant plants (Supplemental Figure S1) were segregated at a ratio of roughly 1:1, and no homozygote was identified (Supplemental Table S1). Pollination of wild-type (WT) plants with pollen from the syp32/+ mutant resulted in approximately 36% of syp32-1/+ (175/[313 + 175]) and 41% of syp32-2/+ (78/[78 + 114]) offspring with the T-DNA insertion (Table 1), indicating that the syp32 mutation was transmitted through the male gametophytes with reduced efficiency. When the stigma of the syp32/+ mutant was pollinated with WT pollen, only 1.2% of syp32-1/+ (6/[6 + 478]) and none of syp32-2/+ (n = 152) progenies bore the insertion (Table 1), suggesting that syp32 mutations cannot be transmitted through the female gametophyte.
No discernable phenotype was observed in the syp31/− mutant (Supplemental Figure S1), raising the question whether SYP31 functions redundantly with SYP32. To answer this question, syp31/+ syp32-1/+ and syp31/+ syp32-2/+ double mutants were generated. Interestingly, no syp31/− syp32/+ plant was recovered in F2 progenies of syp31/+ syp32/+ self-crosses (n = 460). Pollination of emasculated WT flowers with pollen from the syp31/+ syp32/+ mutant resulted in no syp31/+ syp32-1/+ (n = 233) or syp31/+ syp32-2/+ (n = 201) plants (Table 1). Together, the results demonstrated a complete failure in the transmission of syp31 syp32 alleles through the male gametophyte.
Male transmission rate of syp31 syp32 mutations is rescued to largely normal levels by pSYP32:SYP32 but not pSYP32:SYP31 transgenes
To test whether SYP32 can complement the male transmission defect of the double mutants, a construct with a 2-kb promoter region of SYP32 (in relation to the first ATG) driving its cDNA was introduced into the syp31/+ syp32/+ plants. Pollen from syp31/+ syp32-1/+ pSYP32:SYP32/pSYP32:SYP32 and syp31/+ syp32-2/+ pSYP32:SYP32/pSYP32:SYP32 plants was used to fertilize emasculated WT flowers (Table 2). PCR-aided genotyping of the progenies revealed that the male transmission rate of syp31/+ syp32/+ pSYP32:SYP32/pSYP32:SYP32 pollen was increased to above 0.8 (Table 2), suggesting nearly complete rescue of syp31 syp32 male transmission by the SYP32 transgene. To find out the functional differences between SYP31 and SYP32, we introduced SYP31 cDNA under the control of the SYP32 promoter into syp31/+ syp32-1/+ plants to generate the syp31/+ syp32-1/+ pSYP32:SYP31/pSYP32:SYP31 lines. Interestingly, the male transmission efficiency was increased to only ∼0.23 (Table 2). Therefore, SYP31 and SYP32 are partially redundant in pollen development where SYP32’s function is predominant.
| Crosses (Female × Male) . | Progenies . | TEF . | TEM . | |||||
|---|---|---|---|---|---|---|---|---|
| WT . | 31/+ . | 32-1/+ . | 32-2/+ . | 31/+;32-1/+ . | 31/+;32-2/+ . | |||
| +/+ × 31/+ 32-1/+ | 96 | 87 | 50 | 0 | NA | 0 | ||
| +/+ × 31/+ 32-2/+ | 89 | 80 | 32 | 0 | NA | 0 | ||
+/+ ×31/+ 32-1/+ p32:32/p32:32 | 90 | 87 | 79 | 80 | NA | 0.89 | ||
+/+ ×31/+ 32-2/+ p32:32/p32:32 | 69 | 60 | 52 | 58 | NA | 0.84 | ||
+/+ × 31/+ 32-1/+ p32:31/p32:31 (line1) | 88 | 76 | 46 | 23 | NA | 0.26 | ||
+/+ × 31/+ 32-1/+ p32:31/p32:31 (line2) | 86 | 66 | 55 | 16 | NA | 0.19 | ||
| Crosses (Female × Male) . | Progenies . | TEF . | TEM . | |||||
|---|---|---|---|---|---|---|---|---|
| WT . | 31/+ . | 32-1/+ . | 32-2/+ . | 31/+;32-1/+ . | 31/+;32-2/+ . | |||
| +/+ × 31/+ 32-1/+ | 96 | 87 | 50 | 0 | NA | 0 | ||
| +/+ × 31/+ 32-2/+ | 89 | 80 | 32 | 0 | NA | 0 | ||
+/+ ×31/+ 32-1/+ p32:32/p32:32 | 90 | 87 | 79 | 80 | NA | 0.89 | ||
+/+ ×31/+ 32-2/+ p32:32/p32:32 | 69 | 60 | 52 | 58 | NA | 0.84 | ||
+/+ × 31/+ 32-1/+ p32:31/p32:31 (line1) | 88 | 76 | 46 | 23 | NA | 0.26 | ||
+/+ × 31/+ 32-1/+ p32:31/p32:31 (line2) | 86 | 66 | 55 | 16 | NA | 0.19 | ||
For single mutants, TE = (progeny with insertion/progeny without insertion) ×100%; for double mutants: TE = (progeny with both insertions/progeny without insertion) ×100%; TEF and TEM, female and male transmission efficiency, respectively.
NA, not applicable; TE, transmission efficiency.
| Crosses (Female × Male) . | Progenies . | TEF . | TEM . | |||||
|---|---|---|---|---|---|---|---|---|
| WT . | 31/+ . | 32-1/+ . | 32-2/+ . | 31/+;32-1/+ . | 31/+;32-2/+ . | |||
| +/+ × 31/+ 32-1/+ | 96 | 87 | 50 | 0 | NA | 0 | ||
| +/+ × 31/+ 32-2/+ | 89 | 80 | 32 | 0 | NA | 0 | ||
+/+ ×31/+ 32-1/+ p32:32/p32:32 | 90 | 87 | 79 | 80 | NA | 0.89 | ||
+/+ ×31/+ 32-2/+ p32:32/p32:32 | 69 | 60 | 52 | 58 | NA | 0.84 | ||
+/+ × 31/+ 32-1/+ p32:31/p32:31 (line1) | 88 | 76 | 46 | 23 | NA | 0.26 | ||
+/+ × 31/+ 32-1/+ p32:31/p32:31 (line2) | 86 | 66 | 55 | 16 | NA | 0.19 | ||
| Crosses (Female × Male) . | Progenies . | TEF . | TEM . | |||||
|---|---|---|---|---|---|---|---|---|
| WT . | 31/+ . | 32-1/+ . | 32-2/+ . | 31/+;32-1/+ . | 31/+;32-2/+ . | |||
| +/+ × 31/+ 32-1/+ | 96 | 87 | 50 | 0 | NA | 0 | ||
| +/+ × 31/+ 32-2/+ | 89 | 80 | 32 | 0 | NA | 0 | ||
+/+ ×31/+ 32-1/+ p32:32/p32:32 | 90 | 87 | 79 | 80 | NA | 0.89 | ||
+/+ ×31/+ 32-2/+ p32:32/p32:32 | 69 | 60 | 52 | 58 | NA | 0.84 | ||
+/+ × 31/+ 32-1/+ p32:31/p32:31 (line1) | 88 | 76 | 46 | 23 | NA | 0.26 | ||
+/+ × 31/+ 32-1/+ p32:31/p32:31 (line2) | 86 | 66 | 55 | 16 | NA | 0.19 | ||
For single mutants, TE = (progeny with insertion/progeny without insertion) ×100%; for double mutants: TE = (progeny with both insertions/progeny without insertion) ×100%; TEF and TEM, female and male transmission efficiency, respectively.
NA, not applicable; TE, transmission efficiency.
Double mutations in SYP31 and SYP32 cause pollen abortion
The failure of male transmission in syp31/+ syp32/+ double mutants is indicative of defective pollen development. Indeed, about 22% of pollen from the syp31/+ syp32/+ mutant was shriveled and not viable (Figure 1B and D), close to the theoretical value of 25%. By contrast, >99% of pollen from either the WT or single mutants (syp31/−, syp32-1/+, or syp32-2/+) appeared normal (Figure 1B and C). Consistently, shriveled pollen was clearly observed by scanning electron microscopy (SEM), although the exine exhibited a similar pattern to that of the WT (Supplemental Figure S2).
To further confirm that the abortion of pollen was caused by a gametophytic, not a sporophytic defect, quartet (qrt1) mutants were utilized where the four microspores produced by each microsporocyte do not separate from each other throughout pollen development (Preuss et al., 1994). We introduced the syp31 and syp32 alleles into the qrt1 background to generate syp31/+ syp32-1/+qrt1/− and syp31/+ syp32-2/+ qrt1/− plants. Our results showed that zero, one, or two shriveled syp31 syp32 pollens were produced (Figure 1D), which was expected for syp31/+ syp32/+ double mutants where the number of shriveled pollens depended on the segregation of the chromosomes in each meiosis and whether recombination occurred or not. In short, these results supported the conclusion that the aborted pollen grains represented syp31 syp32 gametes.
syp31 syp32 pollen is arrested during the transition from the microspore to the bicellular stage
The consecutive stages of pollen development in Arabidopsis are well defined and can be easily visualized by DAPI staining to follow the number of nuclei (Owen and Makaroff, 1995; Park and Howden, 1998; Backues et al., 2010). At the unicellular microspore (UNM) stage, an average of ∼2.1% pollen from the syp31/+ syp32/+ mutant displayed no DAPI staining, with a similar ratio to that of the WT (1.17%; Figure 2A and B). However, about 18.4% BCP, 19% TCP, and 19.4% MP from the syp31/+ syp32/+ mutant was smaller in size and displayed aberrant or no DAPI staining, at ratios significantly higher than those of the WT (Figure 2A and B). Taken together, the development of syp31 syp32 pollen was arrested during the transition from UNM to BCP. Consistently, endogenous expression of SYP31 and SYP32 was weak in the UNM stage, while the expression dramatically increased throughout the later stages of pollen development, as indicated by β-glucuronidase (GUS) assays performed with pSYP31:GUS and pSYP32:GUS plants (2-kb in relation to ATG for each construct; Figure 2C and D; Supplemental Figure S3).
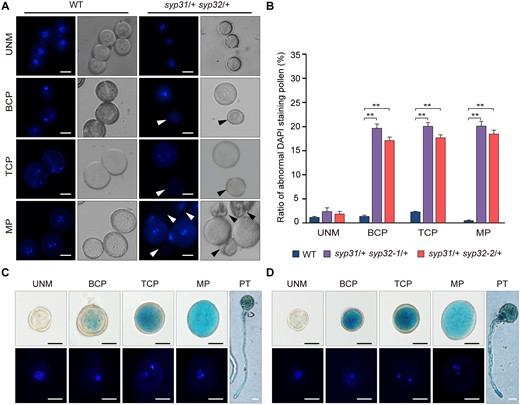
syp31 syp32 pollen is arrested during the transition from the microspore stage to the bicellular stage. A, UNM, BCP, TCP, and MP pollen from WT and syp31/+ syp32/+ anthers were analyzed by DAPI staining. Arrowheads indicate the abnormally stained pollen. B, Statistical analysis of DAPI-stained pollen at different developmental stages (n > 500 pollen from three individual plants was analyzed for each line). Values represent the means SD, **P < 0.01 by two-tailed Student’s t test. C and D, Expression pattern of SYP31 (C) and SYP32 (D) in UNM, BCP, TCP, and MP and pollen tubes (PTs). Nineteen of 22 pSYP31:GUS and 20 of 30 pSYP32:GUS transgenic lines displayed a similar staining pattern. GUS-positive pollen was stained with DAPI before observation. Bars = 10 μm in (A), (C), and (D).
syp31 syp32 pollen is defective in the formation of the PMI cell plate and intine
Aniline blue-stained callose is almost absent in WT pollen at the late bicellular (Figure 3A) and the tricellular stages (Johnson and McCormick, 2001; Backues et al., 2010). However, ∼20% of pollen from syp31/+ syp32-1/+ plants, and ∼15% from syp31/+ syp32-2/+ plants showed ectopic callose deposition (Figure 3A; white arrows), indicating defective cell wall deposition or membrane trafficking (Backues et al., 2010). Callose is not a common component of mature cell walls and is usually only abundant in newly formed cell plates. During PMI, a hemispherical-shaped callosic cell plate is formed to enclose the vegetative nucleus (Figure 3B; Park and Twell, 2001; Park et al., 2004). In syp31/+ syp32/+ anthers, ∼14% of PMI pollen showed abnormal cell plate formation (Figure 3C–F). Among them, various abnormalities, such as pollen with a large callosic aggregate at the cell cortex without the formation of the cell plate (Figure 3C), pollen with a cell wall stub (Figure 3D), and pollen accumulating vesicle-like structures (Figure 3E), were detected.
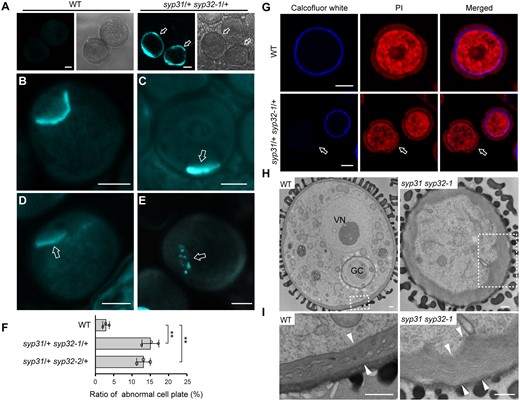
Cell plate and intine defects in syp31 syp32 BCP. A, Ectopic callose deposition in syp31 syp32-1 late BCP. n = 317, n = 265, and n = 204 pollen for WT, syp31/+ syp32-1/+, and syp31/+ syp32-2/+ plants, respectively. Arrows in (A) indicate abnormal pollen. B–E, Aniline blue staining of PMI pollen of WT (B) and syp31 syp32-1 (C–E). Arrows in (C)–(E) indicate abnormal cell plates. F, The statistical analysis of PMI pollen positive for aniline blue staining. n = 202, n = 167, and n = 183 pollen for WT, syp31/+ syp32-1/+ and syp31/+ syp32-2/+ plants, respectively. Pollen from three plants was counted for each genotype. Values represent the means SD, **P < 0.01 by two-tailed Student’s t test. G, Cellulose deposition is defective in syp31 syp32-1 pollen. Cellulose and cytoplasm are labeled with calcofluor white (blue) and PI (red), respectively. Arrows indicate pollen with no Calcofluor white staining. H, TEM analysis revealed abnormal intine deposition in syp31 syp32-1 pollen. GC, generative cell; VN, vegetative nucleus; I, Magnified images of the boxed areas in (H). Intine thickness is indicated by pairs of white arrowheads. Bars = 5 μm (A–E and G) and 500 nm (H and I).
Cellulose is produced by cellulose synthase complexes, which are assembled at the Golgi and then transported to the plasma membrane along the secretory pathway (McFarlane et al., 2014). The intine of WT pollen was strongly stained by the cellulose dye calcofluor white (Figure 3G; Hoedemaekers et al., 2015), while ∼20% of pollen of syp31/+ syp32/+ plants (10/45, syp31/+ syp32-1/+; 11/60, syp31/+ syp32-2/+) was not stained (Figure 3G).
Corroborating these observations (Figure 3A and G), the intine of ∼24% of pollen from syp31/+ syp32-1/+ plants was unevenly deposited, and appeared greater in thickness and lighter in electron density (n = 34; Figure 3H and I), as revealed by transmission electron microscopy (TEM). This phenotype resembled some cellulose-deficient mutant pollen previously reported (Persson et al., 2007; Jozwiak et al., 2015). It has been reported that cellulose deficiency and compromised cell wall integrity frequently go hand in hand with the active synthesis of callose (Lukowitz et al., 2001; Desprez et al., 2002; Gillmor et al., 2005; Van Damme et al., 2006; Persson et al., 2007). Taken together, the double mutations in SYP31 and SYP32 lead to defective cell plate and intine formation, which is most likely caused by improper transport and deposition of cell wall materials.
Golgi-stack integrity is impaired in syp31 syp32 mutant pollen
Since SYP31 and SYP32 are Golgi-localized SNAREs, we analyzed Golgi morphology in syp31 syp32-1 pollen. GFP-Endomembrane Protein12 (EMP12) in WT plant labels a Golgi body as a bar or disk depending on the viewing angle, which reflects typical Golgi morphology (Figure 4A, left-hand graph; Tan et al., 2016). EMP12-labeled Golgi bodies in ∼22% of pollen from the syp31/+ syp32-1/+ mutant appeared much rounder and fuzzier, and the background fluorescence was greatly increased (Figure 4A, middle graph). Occasionally, Golgi bodies were hardly discerned (Figure 4A, right-hand graph). The length/width ratio of the GFP-EMP12 signal of all distinguishable Golgi bodies was measured (Figure 4B and C) where 64% of Golgi bodies with length/width ≥2 were found in WT pollen, but none in mutant pollen (Figure 4B).
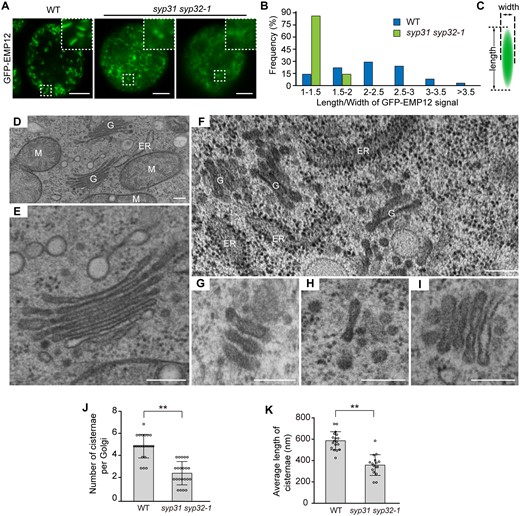
Golgi stack integrity is impaired in syp31 syp32-1 pollen. A, Confocal images of WT and syp31 syp32-1 BCP expressing pUbiquitin10:GFP-EMP12. Insets are 3× magnified images of the selected areas. B, The distribution of the Golgi length/width ratio in BCP. More than 200 Golgi bodies were counted in five WT and syp31 syp32-1 pollens. C, Schematic diagram showing the length and the width of a Golgi body. D–I, TEM images of WT pollen (D and E), syp31 syp32-1 BCP (F–H), and syp31 syp32-1 unicellular pollen (I). G, Golgi apparatus; M, mitochondria. J–K, Statistical analysis of Golgi abnormalities in syp31 syp32-1 BCP. n = 22 Golgi bodies counted for each genotype in (J). n = 18 (WT), n = 16 (syp31 syp32-1) Golgi bodies were counted in (K). Values represent the means SD, **P < 0.01 by two-tailed Student’s t test. Bars = 5 μm in (A), 200 nm in (D–I).
Next, ultrathin sections from high-pressure frozen, freeze-substituted anther samples were prepared for TEM observation. At the bicellular stage, a typical Golgi body in WT pollen displayed its characteristic morphology with approximately five cisternae and an average 580 nm of cisternae length (Figure 4D–E and J–K). In ∼24% of pollen of syp31/+ syp32-1/+ plants, cell wall materials were unevenly deposited, making them readily discernible from the WT (Figure 3H and I). In the aberrant pollen, Golgi morphology disruption was evident in two aspects. First, Golgi bodies lost their characteristic “stacks,” having only 2–3 cisternae left (Figure 4F, G, and J), and in some extreme cases, only one short cisterna left (Figure 4H). These remaining cisternae were surrounded by many vesicle-like structures (Figure 4H). Second, the average cisternae length was dramatically reduced from 580 nm to 361 nm (Figure 4F, G, and K). Moreover, when it was traced back to the unicellular stage, the mutant pollen already displayed a similar, yet milder, Golgi phenotype (Figure 4I and Supplemental Figure S4). Together, these results demonstrate the roles of SYP31 and SYP32 in maintaining Golgi integrity in pollen.
Membrane trafficking is disrupted by syp31 syp32 double mutations
Next, we sought to evaluate the trafficking defects caused by the double mutations. mCherry-HDEL is an ER-localized protein (Nelson et al., 2007); it displayed a reticulate ER pattern in WT pollen (Figure 5A). By contrast, mCherry-HDEL lost its ER pattern completely, and appeared in the vacuole or as punctate structures in syp31 syp32-1 pollen (Figure 5A); therefore, it was not properly retrieved by the ER, but routed to the vacuole. Highly methylesterified pectins labeled by JIM7 antibody are thought to be secreted to the apoplast after synthesis at the Golgi (Li et al., 2019). Unlike in the WT, punctate JIM7-positive structures accumulated in syp31 syp32-1 pollen (Figure 5B). Together with the loss of EMP12 from the periphery of the Golgi (Figure 4A), these findings indicated a disruption of retrograde and anterograde transport by syp31 syp32 double mutations.
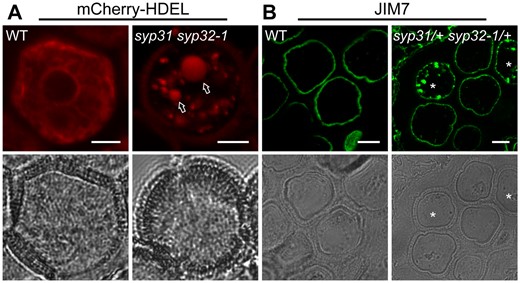
Membrane trafficking is defective in syp31 syp32-1 pollen. A, Confocal images of WT and syp31 syp32-1 unicellular pollen expressing pUbiquitin10: mCherry-HDEL. Vacuole signals (arrows) were detected in ∼19% of pollen of syp31/+ syp32-1/+ plants (n = 79). B, JIM7-labeling of pectins was blocked inside BCP (8/38) of mutants as punctuate structures. Semi-thin anther sections were used for labeling. Asterisks indicate JIM7 positive signals. Bars = 5 μm.
SYP31 and SYP32 directly interact with COG3
To explore the protein interaction network of SYP31 and SYP32, the Arabidopsis cell line (PSB-d) expressing N-terminal protein G/Streptavidin binding peptide (GS)-tagged SYP31 (GS-SYP31), SYP32 (GS-SYP32), or GFP (GS-GFP) fusion proteins were generated. These proteins were purified from Arabidopsis cell extracts using tandem affinity purification (TAP) chromatography (Van Leene et al., 2011), and TAP elutions were analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE; Figure 6A) and liquid chromatography-tandem mass spectrometry (LC-MS/MS; Figure 6B; Supplemental Data Set 1). After the removal of protein G during TAP purification, GS-GFP (∼34 kD), GS-SYP31 (∼45 kD), and GS-SYP32 (∼49 kD) showed up in the TAP elution at the expected molecular weight on the SDS-PAGE (Figure 6A). LC-MS/MS identified a number of proteins that were copurified with GS-SYP31 and GS-SYP32 but not with GS-GFP. Among them, four out of eight subunits of the COG complex (COG1, COG2, COG3, and COG4) and several subunits of the COPI coat were identified (Figure 6B).
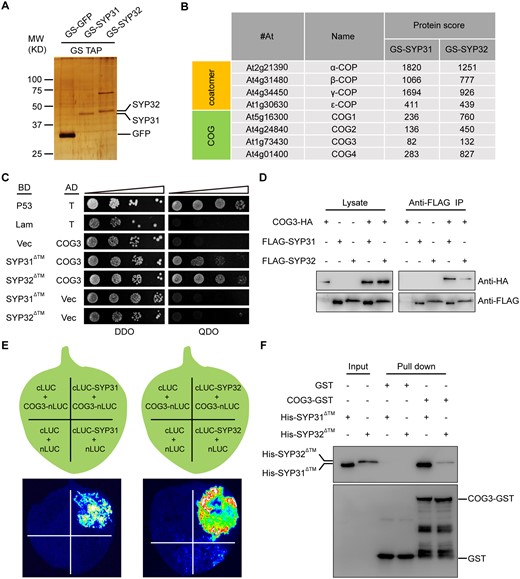
SYP31 and SYP32 interacted with COG3. A, Silver-stained SDS-PAGE gel of TAP-purified GS-GFP, GS-SYP31, and GS-SYP32 elutions. B, Abridged list of proteins that copurified with GS-SYP31 and GS-SYP32 identified by LC-MS/MS. The Mascot protein score is shown. C, SYP31 and SYP32 interacted with COG3 in a Y2H assay. BD-p53 + AD-T served as a positive control. BD-Lam + AD-T, AD-COG3 + BD, BD-SYP31ΔTM/SYP32ΔTM + AD served as negative controls. Columns in each panel represent serial decimal dilutions. DDO, double dropout medium (-Trp –Leu); QDO, quadruple drop out (-Trp –Leu –Ade -His). D, SYP31 and SYP32 co-immunoprecipitated with COG3 in vivo. Lysates of N. benthamiana leaves were tested for protein expression (left panel). The immunoprecipitates were detected using rabbit anti-HA or FLAG antibodies (right panel). E, SYP31 and SYP32 interacted with COG3 in an LCI assay. The Luc signal was captured using a low-light cooled charge-coupled device camera. F, SYP31 and SYP32 interacted with COG3 in a GST pull-down assay. Bound His-SYP31ΔTM or His-SYP32ΔTM was detected using mouse anti-His antibodies. The blot was stripped and reprobed with mouse anti-GST antibodies to ensure the quality and coupling of the bait proteins.
The COG complex is a vesicle tethering complex located at the Golgi and COG3 was previously reported by our lab to have a role in COPI trafficking, Golgi integrity maintenance, and pollen tube growth (Tan et al., 2016). As a continuous effort to study COG function, yeast two-hybrid (Y2H) screening was performed using COG3 as bait. By coincidence, SYP32 was found to be a potential interacting partner (approximately 8-106 yeast transformants were screened, and three positive clones identified corresponded to the N-terminus of SYP32). Therefore, the interactions of SYP31 and SYP32 with COG3 were further explored. SYP31 or SYP32 without the transmembrane domain (SYP31ΔTM and SYP32ΔTM) interacted with COG3 in yeast (Figure 6C). In vivo co-immunoprecipitation (Co-IP) assay showed that COG3-HA was captured by Flag-tagged SYP31 or SYP32 from the N. benthamiana lysates (Figure 6D). In a split luciferase (LUC) complementation assay, COG3-nLUC was able to reconstitute a high luciferase activity with cLUC-SYP31 or cLUC-SYP32 whereas other control pairs of proteins could not (Figure 6E). Furthermore, GST-COG3, but not GST alone, was able to pull down His-SYP31ΔTM and His-SYP32ΔTM in vitro (Figure 6F). Taken together, SYP31 and SYP32 interacted with the COG3 subunit of the COG complex in vitro and in vivo.
SYP31 and SYP32 recruit COG3 to the Golgi apparatus
When transiently expressed in N. benthamiana leaf epidermal cells, COG3 colocalized with SYP31 and SYP32 (Figure 7A), in agreement with previous reports (Uemura et al., 2004; Geldner et al., 2009; Tan et al., 2016; Uemura et al., 2019). We next examined the requirement of SYP3 for the Golgi association of COG3. Remarkably, COG3 remained completely diffused in the cytosol with no Golgi signal in syp31 syp32-1 pollen (Figure 7B). As a control, many Golgi-associated γ-COP signals could still be observed, although with increased background signals (Figure 7C), which is in agreement with the previous reports that ADP-ribosylation factor1 (ARF1) recruits γ-COP to the Golgi bodies (Donaldson et al., 1992; Singh et al., 2018). These data together with the interaction studies (Figure 6) suggested that the Golgi localization of COG3 was achieved by directly interacting with SYP31 and SYP32.

SYP31 and SYP32 colocalize with COG3 and are essential for the Golgi localization of COG3. A, SYP31 or SYP32 colocalize with COG3 when transiently expressed in N. benthamiana leaf epidermal cells. The linear Pearson correlation coefficient (rp) between mCherry-SYP3 and COG3-GFP was obtained using ImageJ with the PSC colocalization plug-in. B, Loss of COG3 Golgi signal in ∼22% of pollen from syp31/+ syp32-1/+ expressing pLAT52:COG3-GFP (n = 128). C, γ-COP localization in ∼24% of pollen from the syp31/+ syp32-1/+ mutant expressing pLAT52:γ-COP- mCherry (n = 83). D, Confocal images of WT and cog3 MP expressing pSYP31:mCherry-SYP31 and pSYP32:mCherry-SYP32. BCP was examined in (B and C). Boxed regions were selected at random and enlarged in the right-hand panels (A) or lower panels (B and C). Bars = 10 μm in (A), 5 μm in (B, C, and D).
On the contrary, Loss of COG3 had no obvious effect on the Golgi localization of SYP31/32, although the background signals were increased as well (Figure 7D).
Discussion
SYP31 and SYP32 function redundantly in male transmission and pollen development
In this study, we explored the physiological roles of SYP3s. We found that pollen viability of syp31/− or syp32/+ single mutants was unaffected (Figure 1), but simultaneous loss of SYP31 and SYP32 caused an arrest in pollen development from the microspore to the bicellular stage (Figures 1 and 2), and led to a complete failure in male transmission (Table 1). pSYP32:SYP32 could restore the male transmission efficiency from 0 to 0.89 in the syp31/+ syp32/+ double mutant (Table 2), whereas the pSYP32:SYP31 transgene only restored it to ∼0.23 (Table 2). Our results suggest that SYP31 and SYP32 play partially redundant roles in pollen development where SYP32 is possibly more important than SYP31. Further study will be necessary to fully elucidate the functional differences between SYP31 and SYP32.
The SYP3–COG interaction provides a part of the molecular mechanisms for severe Golgi disruption and trafficking defects in syp31 syp32 pollen
Our results suggest that the formation of abnormal Golgi structures results from a defect in membrane trafficking. In cog3 pollen, EMP12-positive COPI vesicles lost their tight associations with the periphery of the Golgi bodies, resulting in shortened Golgi with a normal number of cisternae (Tan et al., 2016). The ER pattern labeled by mCherry-HDEL was normal in cog3 pollen (Supplemental Figure S5), suggesting that Golgi-ER retrograde trafficking of mCherry-HDEL-positive COPI vesicles was independent of the COG complex, and this is in agreement with the previous report that retrieval of mCherry-HDEL occurred at the Cis-Golgi (Silva-Alvim et al., 2018). We propose that COG3 might be mainly required for intra-Golgi COPI trafficking. The milder cellular disorders in the cog3 mutant provided a possible explanation for its normal pollen grain development, but not pollen tube growth where secretory activities are more demanding (Tan et al., 2016).
In syp31 syp32 pollen, the secretion blockage of JIM7-positive vesicles from the Golgi/trans-Golgi network to the apoplast (Figure 5) and partial loss of EMP12 from the Golgi (Figure 4) were similarly observed, as in the cog3 mutant (Supplemental Figure S5; Tan et al., 2016). In addition, mCherry-HDEL trafficking between the ER and Golgi was disrupted and it was targeted to the vacuole (Figure 5), consistent with a report suggesting that HDEL promotes vacuolar targeting of proteins that escape the ER (Gomord et al., 1997). As endomembrane trafficking and Golgi integrity are intimately integrated, Golgi disruption was expected to be more severe than in cog3 pollen. Indeed, the Golgi stacks largely fell apart and displayed fewer and shorter cisternae surrounded by accumulating vesicles in syp31 syp32-1 pollen (Figure 4 and Supplemental Figures S4 and S6). The severe disruption of Golgi-associated trafficking and Golgi structure might be the cause of an early abortion of syp31 syp32 pollen at the bicellular stage. Interestingly, in the unicellular pollen, the Golgi phenotype was milder (Compare Figure 4 to Supplemental Figure S4) and pollen development was largely normal, suggesting that Golgi bodies were gradually damaged throughout pollen development.
Nevertheless, it could not be absolutely excluded that SYP3s might work together with other tethering complexes between Golgi cisternae for a tight inter-cisternae association. The detailed molecular mechanism of SYP3s in Golgi needs further study.
SYP31 and SYP32 recruit COG3 to the Golgi apparatus
It has been suggested that the vesicular tethering and fusion in membrane trafficking are orchestrated by direct physical interactions between tethering factors, SNAREs, and Sec1/Munc18 (SM) proteins (Hong and Lev, 2014). In this study, using SYP31 and SYP32 proteins as bait, four COG subunits (COG1-4) of the COG complex were copurified in vivo by LC-MS/MS. Reciprocally, using COG3 as a bait, SYP32 was found to be a binding partner in a Y2H assay. Further analyses verified a direct interaction between COG3 and SYP3 proteins (Figure 6). Remarkably, COG3 completely lost its Golgi association and diffused in the cytosol of syp31 syp32-1 pollen (Figure 7), indicating that COG3 was recruited to the Golgi by SYP3 proteins. Meanwhile, SYP3s still localize at the Golgi to a large extent in the cog3 mutant. This is different from what happens in mammalian cells, where COG complexes were shown to serve as spatial markers for distinct SNARE complexes (Laufman et al., 2013; Willett et al., 2013). In the syp31 syp32 mutant, partial loss of EMP12 in the Golgi is indicative of defective intra-Golgi retrograde trafficking of COPI (Figure 4; Gao et al., 2012). Therefore, the SYP3s-COG3 interaction in plant cells might reflect SYP3s in the intersection of Golgi-related anterograde and retrograde trafficking.
In summary, we demonstrated that SYP31 and SYP32 play essential and partially overlapping roles in membrane trafficking, Golgi structure maintenance, and pollen development.
Materials and methods
Plant materials and growth conditions
Arabidopsis thaliana (L.) Heynh ecotype Columbia-0 (Col-0) was used in the experiments. The T-DNA insertion lines syp31/+ (SALK_057421C), syp32-1/+ (GK-109A09), and syp32-2/+ (GK-920F05) were obtained from the Arabidopsis Biological Resource Center (ABRC). Seeds were surface sterilized for 1 min in 70% (v/v) ethanol, followed by 10-min incubation in 2% (v/v) NaClO with occasional mixing. After washing, seeds were placed on Murashige and Skoog (MS) plates containing 1% (w/v) agar. After 7 d, the seedlings were transferred to soil and grown at 22°C with a 16 h light/8 h dark photoperiod in a growth room. Nicotiana benthamiana plants were grown at 25°C with a 16 h light/8 h dark photoperiod in a growth chamber.
Complementation of syp31/+; syp32/+ mutants
Transgenic plants were generated using the floral dip method (Clough and Bent, 1998). For complementation, the open reading frame of SYP31 or SYP32 was cloned into the pCAMBIA1305 vector under the control of the SYP32 promoter (2,000 bp upstream of the start codon) to generate the pSYP32:SYP31 and pSYP32:SYP32 constructs, which were then introduced into syp31/+ syp32/+ plants. syp31/+ syp32/+ plants homozygous for the pSYP32:SYP31 or pSYP32:SYP32 transgene were selected from the F2 progenies grown on MS medium supplied with 25 mg L−1 hygromycin.
Phenotype characterization
To determine the pollen development stage, anthers from unopened buds were dissected and gently squashed under a coverslip in 4′,6-diamidino-2-phenylindole (DAPI) staining solution (0.1 M sodium phosphate pH 7.0, 1 mM EDTA, 0.1% Triton X-100 [V/V], and 0.5 mg/mL DAPI). The prepared samples were observed under an OLYMPUS BX53 microscope using a DAPI filter set. Pollen viability was examined with Alexander staining as described by Alexander (1969).
Confocal laser scanning microscopy
Observation of various fluorescent fusion proteins, immunolabeling, and cytochemical staining experiments were conducted on a confocal laser scanning microscope (Zeiss, LSM710). GFP signals were excited at 488 nm and emission was detected at 505–530 nm. For propidium iodide (PI) staining and mCherry observation, the excitation wavelength was 561 nm, while the emission wavelength was 575–625 nm. For aniline blue and calcofluor white observation, the settings were adjusted to excitation wavelength 405 nm, emission wavelength 440–500 nm.
Semi-thin sectioning of anthers
Anthers were collected and fixed in a solution (4% paraformaldehyde [w/v], 0.25% glutaraldehyde [v/v], 0.1 M phosphate buffer pH 7.2) at 4°C overnight. After washing 15 min each for 3 times in the phosphate buffer (0.1 M, pH 7.2), the anthers were then dehydrated by a graded series of ethanol (10%, 30%, 50%, 70%, 90%, 95%, and 3 times 100% [v/v]) for about 30 min at each step. Samples were infiltrated in LR White resin according to the following schedule: LR white: 70% ethanol (2:1, v/v), 100% LR white, and 100% LR white (120 min at each step), and left overnight at 4°C after another change of 100% LR white. After being embedded in capsules, polymerization was done at 55°C for 48 h. Semi-thin sections (1 μm) were then prepared using a Leica EM UC7 ultramicrotome.
Immunofluorescence and cytochemical staining
One percent (w/v) calcofluor white and 30 mM PI were added simultaneously to semi-thin sections and incubated for 5 min before washing and observation.
For JIM7 immunofluorescence labeling, semi-thin sections were rehydrated in PBS for 30 min, then incubated in a blocking buffer (2% [w/v] BSA in PBS) for 1 h. After removing the blocking buffer, sections were incubated with JIM7 monoclonal antibodies (1:100 dilution) for 1 h, washed 3 times with PBS, and incubated with Alexa488 conjugated secondary antibodies (1:100 dilution) for another 1 h. After washing 3 times in PBS, the samples were analyzed using a Zeiss LSM710 confocal microscope.
For callose staining, anthers from unopened buds were dissected and gently squashed in 0.1% (w/v) aniline blue under a coverslip before observation.
Scanning and TEM
For SEM observation, pollen grains were mounted on sample stubs. After dehydration in air for 30 min, pollen grains were coated with gold particles (EIKO IB-3). Pollen grains were then observed using a HITACHI S-3000N scanning electron microscope.
Samples for TEM were high-pressure frozen and substituted with 2% (w/v) OsO4 in 100% acetone and infiltrated with Epon resin as described by Otegui et al. (2001). TEM was performed on 90 nm sections using a Hitachi H-7650 transmission electron microscope with a charge-coupled device camera (Hitachi High-Technologies) operating at 80 kV.
TAP purification and mass spectrometry
SYP31, SYP32, or GFP fusions to GS-TAP were cloned into pkngstap using gateway techniques (Van Leene et al., 2011), and transformed into transgenic Arabidopsis PSBd cell lines. TAP of GS-SYP31, GS-SYP32, or GS-GFP was performed as described (Van Leene et al., 2011), with some modifications. Briefly, 6-d-old transgenic PSBd cells expressing GS-SYP31, GS-SYP32, or GS-GFP were lysed using depressurized nitrogen in lysis buffer (20 mM HEPES-KOH pH 7.0, 50 mM KOAc, 1 mM Mg(OAc)2, 250 mM sorbitol, and 1 mM DTT + protease inhibitors). Following lysis, extracts were incubated with 1% (v/v) Triton X-100. After clarification, extracts were incubated with IgG Sepharose 6 Fast Flow beads (GE Healthcare) at 4°C for 1 h, which were then batch washed with wash buffer (10 mM Tris pH 8.0, 150 mM NaCl, 0.1% [v/v] Nonidet P-40, and 5% [v/v] ethylene glycol) 3 times, and incubated with ∼200 units of TEV protease at 22°C for 90 min. The supernatant was collected and incubated with pre-equilibrated Streptavidin Sepharose Resin (GE Healthcare) at 4°C for 1 h. Beads were then batch washed 3 times and eluted with desthiobiotin in wash buffer. Eluants were TCA precipitated and analyzed by SDS-PAGE followed by silver staining or mass spectroscopy analysis. Mass spectrometry and data analysis were performed as described (Mayers et al., 2017).
Protein–protein interacting assays
Y2H analysis was performed using the MATCHMAKER GAL4 Two-Hybrid System (Clontech) according to the manufacturer’s instructions. For protein pair analyses, the cDNAs were cloned into pGBKT7 or pGADT7 vectors (Clontech), and plasmids of each pair were cotransformed into the yeast strain AH109. Transformants were selected on DDO medium at 30°C, while the selection of interactions was conducted on QDO medium at 30°C for 3 d. The experiments were performed at least 3 times independently, and similar results were obtained.
The immunoprecipitation assay was performed as described (Wu et al., 2013), with some modifications. Briefly, Agrobacterium tumefaciens (EHA105) containing p35S:3×FLAG-SYP31, p35S:3×FLAG-SYP32, and p35S:3×HA-COG3 were grown until O.D.600 for around 1.0–1.5. After 48 h of infiltration, the leaves were ground in liquid nitrogen and resuspended in lysis buffer (as described in the TAP experiment above). The total extract was incubated with 1% (v/v) Triton X-100 for 1 h and centrifuged at 16,000 g at 4°C for 30 min, and the supernatant was kept immobilized on anti-FLAG M2 agarose (Sigma A2220) by rotating at 4°C for 2 h. After being washed 5 times, the samples were subjected to SDS–PAGE and a western blot assay with rabbit anti-HA and anti-FLAG antibodies.
Luciferase complementation imaging (LCI) assays and in vitro GST pull-down assays were performed as described (Wu et al., 2017).
All primers used in this study were listed in Supplemental Table S2.
Accession numbers
The accession numbers for DNA sequences used in this study are as follows: SYP31, AT5G05760; SYP32, AT3G24350; COG3, AT1G73430; γ-COP, AT4G34450; EMP12, AT1G10950.
Supplemental data
The following materials are available in the online version of this article.
Supplemental Figure S1. Genotyping of syp31/−, syp32-1/,+ and syp32-2/+ mutants.
Supplemental Figure S2. SEM analysis of MP from WT, syp31/+ syp32-1/+, and syp31/+ syp32-2/+ plants.
Supplemental Figure S3. Expression pattern of SYP31 and SYP32.
Supplemental Figure S4. TEM analysis of WT and syp31 syp32-1 pollen at the unicellular stage.
Supplemental Figure S5. Analyses of trafficking defects in cog3 pollen/pollen tubes.
Supplemental Figure S6. Diagrams of the Golgi apparatus in WT, cog3 and syp31 syp32 pollen.
Supplemental Table S1. Segregation analysis of syp32 alleles.
Supplemental Table S2. The list of primers used in this study.
Supplemental Data Set 1. Detailed LC-MS/MS datasheet of proteins identified with GS-TAP.
Acknowledgments
We thank Ben August and Grzegorz Sabat (University of Wisconsin-Madison) for help with TEM and mass spectroscopy.
Funding
This research was supported by a grant from the National Science Foundation of China (No. 31770202 and No. 32070190), and a grant from the Ministry of Agriculture of China for Transgenic Research (No. 2018ZX08009-20B).
Conflict of interest statement. The authors have declared that no competing interests exist.
Q.R., S.Y.B., and Y.B. conceived the study and designed the experiments. Q.R. performed most of the experiments. X.T., F.L., Y.L., X.L., B.L., J.W., H.Y., L.Q., T.L., S.F., R.G., and W.W. provided technical assistance to Q.R. Q.R., S.Y.B., and Y.B. analyzed data. Q.R. and Y.B. wrote the article. All authors read and approved the manuscript.
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (https://dbpia.nl.go.kr/plphys/pages/general-instructions) is: Yiqun Bao ([email protected]).
References
Author notes
Senior author.



