-
PDF
- Split View
-
Views
-
Cite
Cite
Santiago Cabezas-Camarero, Vanesa García-Barberán, Rebeca Pérez-Alfayate, María Elena Gómez del Pulgar, Maria Nieves Cabrera-Martin, Isabel Casado-Fariñas, Pedro Pérez-Segura, Plasma ctDNA Monitoring of a PTCH1-Mutant Metastatic Adult Medulloblastoma Showing a Durable Benefit With Vismodegib, The Oncologist, Volume 29, Issue 5, May 2024, Pages 377–383, https://doi.org/10.1093/oncolo/oyae026
Close - Share Icon Share
Abstract
Adult medulloblastoma (MB) is a rare disease affecting 0.6 persons per million adults over 19 years of age. The SHH-activated/TP53-wild type is the most common subtype, accounting for 60% of adult MBs, being characterized by mutations in PTCH1, SMO, or the TERT promoter. Several small studies demonstrate objective but short-lived responses to SMO inhibitors such as vismodegib or sonidegib. Like other oncogene-addicted solid tumors, detection of the corresponding drivers through liquid biopsy could aid in the molecular diagnosis and monitoring of the disease through less invasive procedures. However, most studies have only evaluated cerebrospinal fluid as the ctDNA reservoir, and very limited evidence exists on the role of liquid biopsy in plasma in patients with primary central nervous system tumors, including MB. We present the case of a 26-year-old patient with a recurrent MB, in which next-generation sequencing (FoundationOne CDx) revealed a mutation in PTCH1, allowing the patient to be treated with vismodegib in second line, resulting in a durable benefit lasting for 1 year. Using an in-house digital PCR probe, the PTCH1 mutation could be tracked in ctDNA during treatment with first-line chemotherapy and while on treatment with vismodegib, demonstrating a precise correlation with the radiological and clinical behavior of the disease.
SHH-activated/TP53-wild type is the most common subtype of medulloblastoma (MB) in adults, being characterized by PTCH1 loss of heterozygosity in most cases.
Several small studies have demonstrated that SHH-activated/TP53-wild type MB may benefit from SMO inhibitors although responses are usually short lived.
A patient with long-lasting benefit with vismodegib is presented, demonstrating the feasibility of PTCH1-mutation tracking in plasma.
Plasma ctDNA mirrored the clinical and radiological response during treatment with first-line chemotherapy and second-line vismodegib.
Introduction
Medulloblastoma (MB), an embryonal tumor developing in the posterior fossa, is the most common brain malignancy in children but is rare in adults, with a reported incidence of 0.6 cases per 1 million adults older than 19 years of age.1 The SHH-activated/TP53-wild-type subtype, is the most common in adults, accounting for 60% of the cases, and has an intermediate prognosis amongst the 4 currently established MB subgroups, with a 5-year overall survival of 76%.1 It is characterized by mutations in PTCH1, SMO, DDX3X, and TERT promoter in adults, and by mutations in SUFU in pediatric MB.1,2 SMO pathway inhibitors, such as vismodegib and sonidegib have been tested in recurrent SHH-activated MB due to PTCH1 mutations, with notable although short-lived responses.3-10
A few studies using liquid biopsy in patients with MB have been published in recent years, demonstrating the feasibility of ctDNA detection in cerebrospinal fluid (CSF) in approximately half of patients, and allowing for molecular subgroup classification and disease monitoring including minimal residual disease (MRD) detection, and molecular characterization at relapse.11-17 Most studies were conducted using CSF in patients with CNS-only disease, and in the few studies including plasma, ctDNA detection rate was absent or very low and with very low mutation allele frequencies (MAFs). Therefore, the generalizability of liquid biopsy in clinical practice is currently limited due to the risks and discomfort for patients of CSF collection procedures and due to its low sensitivity in plasma.11 However, in patients with MB and extraneural dissemination, liquid biopsy in plasma could be a useful diagnostic and monitoring tool.18
We present the case of a patient with a recurrent MB, showing a 1-year-long benefit with second-line vismodegib, where serial ctDNA PTCH1-mutation detection in plasma allowed for accurate disease monitoring.
Patient Story
A 23-year-old woman was diagnosed in October 2017 with a desmoplatic/nodular type MB of the right cerebellar hemisphere. Biopsy revealed a neoplasm characterized by small blue cells arranged in pale nodular areas with neuronal differentiation (synaptophysin-positive) surrounded by peripheral reticulin-rich areas with densely packed hyperchromatic cells with high proliferative activity. A complete resection was performed followed by adjuvant total craniospinal irradiation with tomotherapy. Since there were no risk factors detected (complete surgical resection with no CSF dissemination), adjuvant chemotherapy was not administered.
Two years later, in September 2019, an 18F-FDG PET-CT scan revealed multiple osteoblastic metastases in the whole axial and appendicular skeleton. Bone marrow biopsy demonstrated infiltration by a malignant neoplasm of small cells growing in nets and rows, surrounded by fibrosis, and with an immunohistochemical profile compatible with MB (CD56 positive and CKAE1/AE3, CD45, myeloperoxidase, and S-100 protein negative). Treatment with carboplatin-etoposide every 3 weeks for 3 cycles followed by a fourth cycle with cyclophosphamide-carboplatin-etoposide (CCE), led to a major partial response at almost all metastatic sites. After suffering from progressive disease in early April 2020, 3 additional cycles of CCE were administered, leading to a major partial response. In October 2020, progression of bone metastases occurred, and next-generation sequencing of the primary tumor revealed a pathogenic mutation in PTCH1 (Y873*; Figure 1). Treatment with the oral SMO inhibitor vismodegib 150 mg daily in 28-day cycles combined with zoledronic acid 4 mg IV every 4 weeks was started, with partial response after 8 weeks, followed by sustained partial response until September 2021 when progressive disease with symptomatic deterioration (bone pain) occurred. Vismodegib was stopped with mild pain improvement. It was reintroduced 6 weeks later and maintained until January 2022, when symptomatic deterioration (bone pain) due to progressive disease occurred. Treatment with vismodegib was well tolerated with alopecia G2 and anemia G2 as the major toxicities.
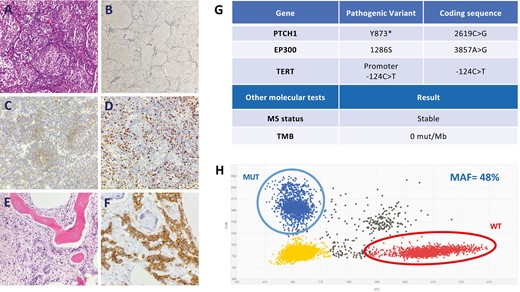
Pathology findings, next-generation sequencing results, and digital PCR analyses of the primary tumor. (A) Desmoplastic/nodular MB with nodular pattern (pale islands surrounded by densely packed poorly differentiated cells; H&E, 200×). (B) The presence of peripheral reticulin-rich areas surrounding well-developed reticulin-poor nodules with neuronal differentiation is one of the most diagnostically helpful features for the desmoplastic/nodular MB variant (Reticulin, 100×). (C) Neuronal differentiation shown by moderate immunoreactivity for sinaptophysin in the pale nodes (Sinaptophysin, 200×). (D) Ki-67 antibody staining shows that proliferative activity predominates in the highly cellular internodular areas (Ki-67, 200×). Bone marrow infiltrated by poorly differentiated cells (E, H&E, 100×) which are CD56-positive, corresponding to metastasis of MB (F, CD56, 400×). (G) The findings of the molecular NGS study (Foundation One CDx) of the primary tumor are shown. (H) Results of the digital PCR of the PTCH1 Y873* mutation in the primary tumor. Encircled left upper dots indicate the mutant fraction. Encircled right lower dots indicate the wild-type fraction.
Since the start of first-line chemotherapy until the end of second-line therapy with vismodegib, blood was serially collected to monitor the PTCH1 somatic mutation in ctDNA. As shown in Fig. 2, 3 weeks after the start of chemotherapy, ctDNA levels were notably decreased becoming undetectable after the second cycle. After the start of vismodegib, the PTCH1 mutation became undetectable in October 2020 until June 2021, consistent with the radiological responses observed on 18F-FDG PET/TC scans. A brisk increase in ctDNA levels occurred in October 2021, in parallel with the radiological progression observed on 18F-FDG PET/CT.
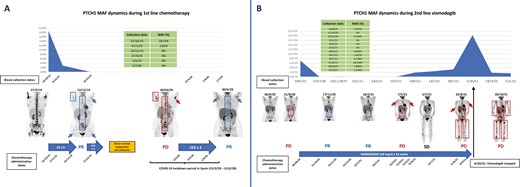
ctDNA PTCH1 mutation dynamics during treatment with first-line platinum-based chemotherapy and second-line vismodegib. (A) The figure shows clearance of the PTCH1 mutation in plasma after 3 cycles of 3-weekly carboplatin-etoposide, being concordant with the near-complete response on the 18F-FDG PET/TC scan from December 2019. Treatment was temporarily stopped in late January 2020 to allow for the collection of progenitor bone marrow cells for a future treatment with high-dose chemotherapy followed by an autologous bone marrow transplant. However, due to the advent of the COVID-19 pandemic, such treatment plan was abandoned to avoid the risks of severe immunosuppression. Tumor progression occurred in March 2023, and treatment with cyclophosphamide-carboplatin-etoposide was restarted in early April 2020, and administered for 3 cycles, leading to a major partial response. PTCH1 ctDNA was negative at 3 different time points during this period, possibly due to the tumor responding to chemotherapy. For more details, please refer to the main text. (B) The figure shows clearance of the PTCH1 mutation in plasma after 3 weeks of treatment with vismodegib and being detectable again at very low MAF in November 2020 and then clearing again until June 2021, being concordant with the near-complete response seen on the 18F-FDG PET/TC scan between November 2020 and February 2021. ctDNA levels started rising in June 2021 and peaked in October 2021, mirroring the radiological disease-progression observed in May, August, and October 2021. Interestingly a dramatic decrease in PTCH1 ctDNA occurred after stopping treatment with vismodegib, that paralleled the symptomatic improvement of the patient, possibly reflecting decreased treatment pressure, although this only lasted for a couple of weeks. Unfortunately, no subsequent blood collections were performed to monitor PTCH1 ctDNA when vismodegib was rechallenged between November 2021 and January 2022 with initial symptomatic improvement but no radiological response. Abbreviation: MAF: mutant-allele frequency. Arrows and boxes in blue tumor lesions that respond or progress as indicated in the figure..
Molecular Tumor Board
Genotyping Studies
In February 2020, a next-generation sequencing multigene panel (FoundationOne CDx) was used for molecular analysis of the primary tumor biopsy.19 The study, among other findings summarized in Figure 1, revealed a pathogenic mutation in PTCH1 (Y873*). To serially monitor the PTCH1 mutation detected in the primary tumor by means of liquid biopsy, 30 mL of peripheral blood were collected at 13 different time points between October 2019 and November 2021. An in-house PCR probe was designed for the study of the PTCH1 mutation (NM_000264.5(PTCH1): c.2619C>G (p.Tyr873Ter)). The QIAmp Circulating Nucleic Acide (QIAGEN, Germany) was used for DNA extraction from 3 mL of plasma from each collected sample. The QuantStudio 3D Digital PCR System (ThermoFisher Scientific, Waltham, MA, USA) was used for digital PCR analysis in ctDNA (Figure 1).
Finally, although no familial history of cancer was reported, an in-house germline study in peripheral leukocyte DNA using digital PCR, was negative for the mutation in PTCH1 indicating the sporadic nature of the patient’s MB (not shown).
The patient gave consent to participate in the study through the signature of an informed consent form. The study was approved by the institutional review board of Hospital Clinico Universitario San Carlos.
Potential Strategies to Target the Pathway and Implications for Clinical Practice
SMO inhibitors such as vismodegib or sonidegib have demonstrated notable objective response rates (ORR) in SHH-activated MB, although they are generally short lived. In 2009, Rudin et al3 reported for the first time a major response with vismodegib in a recurrent MB with extensive bone and soft-tissue metastases, although it only lasted for 3 months. Petrirena et al20 reported a 9-month partial metabolic response with second-line vismodegib in an extraneurally relapsing MB. Kian et al9 recently reported the case of a 24-year-old woman with axillary recurrence who experienced a complete response with first-line vismodegib. Robinson et al7 in 2 phase II studies of patients with recurrent MB, 31 adults (PBTC-025B) and 12 pediatric (PBTC-032) patients, reported 3/31 and 1/12 protocol-specified responses (complete or partial response maintained for at least 8 weeks), with longer progression-free survival in those with SHH-activated MB, and prolonged disease stabilization in 41% of SHH-activated MB. Tumors with PTCH1 loss of heterozygosity were associated with increased duration of therapy with vismodegib. Finally, in a meta-analysis of 5 phases I-II studies, the pooled ORR was 37% in SHH-activated MBs while it was 0 in non-SHH MBs.21
Unfortunately, there is limited understanding of the mechanisms of primary or acquired resistance to hedgehog pathway inhibitors. In the PBTC-025B and PBTC-032 studies, mutations were identified downstream of SMO in 4 non-responding tumors -2 in each group. Three of them harbored somatic mutations in SUFU and a fourth pediatric tumor had a GLI2 amplification, all known to confer resistance to SMO inhibitors in mouse models. The 2 adult tumors harboring SUFU mutations also showed mutations in the PIK3 pathway, including mutations in PIK3CA and PTEN, and one TP53 mutation, the latter known to confer a worse prognosis in SHH-activated MB.1,7 Petrirena et al20 reported acquired mutations in SMO (G477L and L412P) and PIK3CA (H1047A and H1065L) after therapy with vismodegib and sonidegib. On the other hand, in 2 adult and 1 pediatric MB that initially demonstrated response to vismodegib in the PBTC-025B and PBTC-032 trials, tumor biopsies obtained after therapy were devoid of SMO mutations, but harbored mutations in BCOR, BRPF1, NRAS, and SYNE1 in a 16-month responding tumor and in PTEN and GNAS in the other, short-lived responding tumors. Table 1 summarizes these and other studies with SMO inhibitors conducted in patients with adult MB.22
Summary of most relevant studies with SMO inhibitors in patients with SHH-activated adult MB.
| Author (year) . | Study Type . | N . | Agent . | ORR . | PFS . | OS . |
|---|---|---|---|---|---|---|
| Rudin et al. (2009) [3] | Case report | 1 | Vismodegib | Major response | 3 m | DOD 5 m after starting vismodegib |
| LoRusso et al. (2011) [4] | Case report | 1 | Vismodegib | Unconfirmed partial response | — | — |
| Gajjar et al. (2013) [5] | Phase I | 7 | Vismodegib | 1/3 (33%) | — | — |
| Asklund et al. (2013) [22] | Case report | 1 | Vismodegib | Partial metabolic response | ≥5 m | ≥5 m |
| Rodon et al. (2014) [6] | Phase I | 9 | Sonidegib | 3/9 (33%) | — | — |
| Robinson et al. (2015) [7] | Phase II (PBTC-025B: adult pts) (PBTC-032: pediatric pts) | Adult pts: 20 Pediatric pts: 12 | Vismodegib | Adult pts: 3/20 (15%) Pediatric pts: 1/12 (8.3%) | >2 m | — |
| Kieran et al. (2017) [8] | Phase I/II | 10 | Sonidegib | 5/10 (50%) | — | — |
| Petrirena et al. (2018) [20] | Case report | 1 | Vismodegib | Parial metabolic response | 9 m | 27 m |
| Kian et al. (2020) [9] | Case report | 1 | Vismodegib | Complete response | — | Alive 15 m after starting vismodegib |
| Frappaz et al. (2021) [10] | Phase I/II | (A) N = 10 (B) N = 5 (C) N = 9 | (A) Vismodegib + TMZ (B) TMZ (C) Vismodegib | (A) 40% (B) 20% (C) 22% | (A) PFS-6 m: 20% (B) — (C) 237.5% | — |
| Current study | Case report | 1 | Vismodegib | Partial response | 6 m | Died of disease 34 m after starting vismodegib |
| Author (year) . | Study Type . | N . | Agent . | ORR . | PFS . | OS . |
|---|---|---|---|---|---|---|
| Rudin et al. (2009) [3] | Case report | 1 | Vismodegib | Major response | 3 m | DOD 5 m after starting vismodegib |
| LoRusso et al. (2011) [4] | Case report | 1 | Vismodegib | Unconfirmed partial response | — | — |
| Gajjar et al. (2013) [5] | Phase I | 7 | Vismodegib | 1/3 (33%) | — | — |
| Asklund et al. (2013) [22] | Case report | 1 | Vismodegib | Partial metabolic response | ≥5 m | ≥5 m |
| Rodon et al. (2014) [6] | Phase I | 9 | Sonidegib | 3/9 (33%) | — | — |
| Robinson et al. (2015) [7] | Phase II (PBTC-025B: adult pts) (PBTC-032: pediatric pts) | Adult pts: 20 Pediatric pts: 12 | Vismodegib | Adult pts: 3/20 (15%) Pediatric pts: 1/12 (8.3%) | >2 m | — |
| Kieran et al. (2017) [8] | Phase I/II | 10 | Sonidegib | 5/10 (50%) | — | — |
| Petrirena et al. (2018) [20] | Case report | 1 | Vismodegib | Parial metabolic response | 9 m | 27 m |
| Kian et al. (2020) [9] | Case report | 1 | Vismodegib | Complete response | — | Alive 15 m after starting vismodegib |
| Frappaz et al. (2021) [10] | Phase I/II | (A) N = 10 (B) N = 5 (C) N = 9 | (A) Vismodegib + TMZ (B) TMZ (C) Vismodegib | (A) 40% (B) 20% (C) 22% | (A) PFS-6 m: 20% (B) — (C) 237.5% | — |
| Current study | Case report | 1 | Vismodegib | Partial response | 6 m | Died of disease 34 m after starting vismodegib |
Abbreviations: m: months, N: number of patients, ORR: objective response rate, OS: overall survival since the start of SMO inhibitors, PFS: progression-free survival since the start of SMO inhibitors, pts: patients, TMZ: temozolomide.
Summary of most relevant studies with SMO inhibitors in patients with SHH-activated adult MB.
| Author (year) . | Study Type . | N . | Agent . | ORR . | PFS . | OS . |
|---|---|---|---|---|---|---|
| Rudin et al. (2009) [3] | Case report | 1 | Vismodegib | Major response | 3 m | DOD 5 m after starting vismodegib |
| LoRusso et al. (2011) [4] | Case report | 1 | Vismodegib | Unconfirmed partial response | — | — |
| Gajjar et al. (2013) [5] | Phase I | 7 | Vismodegib | 1/3 (33%) | — | — |
| Asklund et al. (2013) [22] | Case report | 1 | Vismodegib | Partial metabolic response | ≥5 m | ≥5 m |
| Rodon et al. (2014) [6] | Phase I | 9 | Sonidegib | 3/9 (33%) | — | — |
| Robinson et al. (2015) [7] | Phase II (PBTC-025B: adult pts) (PBTC-032: pediatric pts) | Adult pts: 20 Pediatric pts: 12 | Vismodegib | Adult pts: 3/20 (15%) Pediatric pts: 1/12 (8.3%) | >2 m | — |
| Kieran et al. (2017) [8] | Phase I/II | 10 | Sonidegib | 5/10 (50%) | — | — |
| Petrirena et al. (2018) [20] | Case report | 1 | Vismodegib | Parial metabolic response | 9 m | 27 m |
| Kian et al. (2020) [9] | Case report | 1 | Vismodegib | Complete response | — | Alive 15 m after starting vismodegib |
| Frappaz et al. (2021) [10] | Phase I/II | (A) N = 10 (B) N = 5 (C) N = 9 | (A) Vismodegib + TMZ (B) TMZ (C) Vismodegib | (A) 40% (B) 20% (C) 22% | (A) PFS-6 m: 20% (B) — (C) 237.5% | — |
| Current study | Case report | 1 | Vismodegib | Partial response | 6 m | Died of disease 34 m after starting vismodegib |
| Author (year) . | Study Type . | N . | Agent . | ORR . | PFS . | OS . |
|---|---|---|---|---|---|---|
| Rudin et al. (2009) [3] | Case report | 1 | Vismodegib | Major response | 3 m | DOD 5 m after starting vismodegib |
| LoRusso et al. (2011) [4] | Case report | 1 | Vismodegib | Unconfirmed partial response | — | — |
| Gajjar et al. (2013) [5] | Phase I | 7 | Vismodegib | 1/3 (33%) | — | — |
| Asklund et al. (2013) [22] | Case report | 1 | Vismodegib | Partial metabolic response | ≥5 m | ≥5 m |
| Rodon et al. (2014) [6] | Phase I | 9 | Sonidegib | 3/9 (33%) | — | — |
| Robinson et al. (2015) [7] | Phase II (PBTC-025B: adult pts) (PBTC-032: pediatric pts) | Adult pts: 20 Pediatric pts: 12 | Vismodegib | Adult pts: 3/20 (15%) Pediatric pts: 1/12 (8.3%) | >2 m | — |
| Kieran et al. (2017) [8] | Phase I/II | 10 | Sonidegib | 5/10 (50%) | — | — |
| Petrirena et al. (2018) [20] | Case report | 1 | Vismodegib | Parial metabolic response | 9 m | 27 m |
| Kian et al. (2020) [9] | Case report | 1 | Vismodegib | Complete response | — | Alive 15 m after starting vismodegib |
| Frappaz et al. (2021) [10] | Phase I/II | (A) N = 10 (B) N = 5 (C) N = 9 | (A) Vismodegib + TMZ (B) TMZ (C) Vismodegib | (A) 40% (B) 20% (C) 22% | (A) PFS-6 m: 20% (B) — (C) 237.5% | — |
| Current study | Case report | 1 | Vismodegib | Partial response | 6 m | Died of disease 34 m after starting vismodegib |
Abbreviations: m: months, N: number of patients, ORR: objective response rate, OS: overall survival since the start of SMO inhibitors, PFS: progression-free survival since the start of SMO inhibitors, pts: patients, TMZ: temozolomide.
There are no approved targeted drugs that may allow for bypassing or reversing resistance to SMO inhibitors in SHH-MB. Interestingly, in basal cell carcinoma (BCC), well-known to be an SHH-activated tumor, taladegib has been shown to inhibit SMO in its wild-type and D473-mutant forms—the latter known to confer resistance to conventional SMO inhibitors.23
As mentioned earlier, PIK3CA mutations occur frequently in SMO-inhibitor-resistant SHH-activated MB and could be a potential target for PIK3CA inhibitors.24-26 Likewise, IDH mutations have been described in SHH-activated MB, and could serve as a potential target for newly-developed IDH-inhibitors such as vorasidenib.27,28
Immune checkpoint inhibitors have been tested in a phase II study of adult patients with pediatric tumors, including patients with MB (11.6%), treated with ipilimumab + nivolumab, with modest benefit.29 Interestingly, SMO inhibitors have been shown to upregulate MHC-I/II and to increase CD8+ T cells in BCC, providing a rationale for evaluating SMO inhibitor and immunotherapy combinations.23
Liquid biopsy in patients with MB has mainly been studied in patients in pediatric care and using CSF as the ctDNA reservoir, with detection rates of approximately 50% of cases in most studies, demonstrating a high accuracy in detecting MRD and genomic evolution characterization.11-17 Very limited evidence exists on the role of ctDNA in plasma in patients with MB. Escudero et al11 reported detection of ctDNA in plasma in 7.7% (1/13 patients) compared to 76.9% (10/13) in CSF. Using digital PCR, Arthur et al17 detected ctDNA in plasma in 4/9 patients compared to 7/12 patients in CSF. Pagès et al16 reported that copy number and other alterations in plasma were detected in 3/230 (1.3%) and 2/74 (2.7%), respectively, compared to 9/46 (20%) and 3/10 (30%), respectively, in CSF. Table 2 summarizes the most relevant studies on liquid biopsy in patients with MB.
| Author (year) . | N . | Reservoir . | No. of samples . | Sequencing method . | Genomic studies . | Detection rate . | Other findings and potential applications . |
|---|---|---|---|---|---|---|---|
| Escudero et al. (2020) [11] | 13 | CSF and Plasma | Serial | Tumor: NGS CSF: NGS and ddPCR Plasma: ddPCR | WES | CSF: 10/13 (76.9%) (baseline) Plasma: 1/13 (7.7%) | CSF ctDNA reflects tumor genomic alterations and evolution, allows MB subgroup classification, and identifies MRD |
| Li et al. (2020) [12] | 3 | CSF | Serial | WGBS and CMS-IP-Seq | WGMeth (WGBS and CMS-IP-Seq) | ctDNA WGMeth precedes positive cytology and identifies markers with predictive and prognostic value | |
| Liu et al. (2021) [13] | 123 | CSF | Serial | NGS | WGS | MRD: 67/105 (64%) MRD SHH vs WNT vs G3 vs G4: 32% vs 63% vs 77% vs 75% Metastatic: 29/34 (85%) Non-metastatic: 38/71 (54%) | High concordance tumor-CSF MRD detection preceded relapse by ≥3 m |
| Sun et al. (2021) [14] | 58 | CSF: 58 Plasma: 6 | Single | NGS | WES | 15/58 (26%) | ctDNA-positive CSF associated with worse prognosis |
| Lee et al. (2022) [15] | 40 | CSF | Single | RNA-seq and HRMS | RNA-seq and HRMS | — | Different transcriptomic profile, cirRNA and metabolic profiles in CSF between pts with MB and pts without MB |
| Pagès et al. (2022) [16] | 258 | CSF Plasma | Single | ULP-WGS | ULP-WGS | CSF CNA: 9/46 (20%) CSF alterations: 3/10 (30%) Plasma CNA: 3/230 (1.3%) Plasma alterations: 2/74 (2.7%) | CSF is more reliable than plasma for genomic analyses in children with MB |
| Arthur et al. (2023) [17] | 12 | CSF and Plasma | Serial | Tumor: NGS CSF: ddPCR | SNCAIP, CTNNB1, PIK3CA, FOSL2, ZNF536, HEXB, MAX, PTCH1, B4GALT1, KDM5D, CDK6, SAMD8, PRMT7, KMT2D, OFD1, SMO, KDM6A | CSF: 7/12 Plasma: 4/9 | CSF ctDNA positive in tumors growing into a CSF reservoir |
| Current study | 1 | Plasma | Serial | dPCR | PTCH1 Y873* | 10/18 timepoints positive 8/8 false negative | ctDNA PTCH1 mutation dynamics mirrored accurately radiological response |
| Author (year) . | N . | Reservoir . | No. of samples . | Sequencing method . | Genomic studies . | Detection rate . | Other findings and potential applications . |
|---|---|---|---|---|---|---|---|
| Escudero et al. (2020) [11] | 13 | CSF and Plasma | Serial | Tumor: NGS CSF: NGS and ddPCR Plasma: ddPCR | WES | CSF: 10/13 (76.9%) (baseline) Plasma: 1/13 (7.7%) | CSF ctDNA reflects tumor genomic alterations and evolution, allows MB subgroup classification, and identifies MRD |
| Li et al. (2020) [12] | 3 | CSF | Serial | WGBS and CMS-IP-Seq | WGMeth (WGBS and CMS-IP-Seq) | ctDNA WGMeth precedes positive cytology and identifies markers with predictive and prognostic value | |
| Liu et al. (2021) [13] | 123 | CSF | Serial | NGS | WGS | MRD: 67/105 (64%) MRD SHH vs WNT vs G3 vs G4: 32% vs 63% vs 77% vs 75% Metastatic: 29/34 (85%) Non-metastatic: 38/71 (54%) | High concordance tumor-CSF MRD detection preceded relapse by ≥3 m |
| Sun et al. (2021) [14] | 58 | CSF: 58 Plasma: 6 | Single | NGS | WES | 15/58 (26%) | ctDNA-positive CSF associated with worse prognosis |
| Lee et al. (2022) [15] | 40 | CSF | Single | RNA-seq and HRMS | RNA-seq and HRMS | — | Different transcriptomic profile, cirRNA and metabolic profiles in CSF between pts with MB and pts without MB |
| Pagès et al. (2022) [16] | 258 | CSF Plasma | Single | ULP-WGS | ULP-WGS | CSF CNA: 9/46 (20%) CSF alterations: 3/10 (30%) Plasma CNA: 3/230 (1.3%) Plasma alterations: 2/74 (2.7%) | CSF is more reliable than plasma for genomic analyses in children with MB |
| Arthur et al. (2023) [17] | 12 | CSF and Plasma | Serial | Tumor: NGS CSF: ddPCR | SNCAIP, CTNNB1, PIK3CA, FOSL2, ZNF536, HEXB, MAX, PTCH1, B4GALT1, KDM5D, CDK6, SAMD8, PRMT7, KMT2D, OFD1, SMO, KDM6A | CSF: 7/12 Plasma: 4/9 | CSF ctDNA positive in tumors growing into a CSF reservoir |
| Current study | 1 | Plasma | Serial | dPCR | PTCH1 Y873* | 10/18 timepoints positive 8/8 false negative | ctDNA PTCH1 mutation dynamics mirrored accurately radiological response |
Abbreviations: CNA: copy number alterations, CMS-IP-Seq: anti-cytosine-5-methylenesulfonate immunoprecipitation sequencing, CSF: cerebrospinal fluid, ctDNA: circulating tumor DNA, dPCR: digital PCR, ddPCR: droplet digital PCR, G3: group 3 MB, G4: group 4 MB, HRMS: high-resolution mass spectrometry, MB: medulloblastoma, MRD: minimal residual disease, NGS: next-generation sequencing, SHH: sonic hedgehog activated MB, ULP-WGS: ultra-low pass whole-genome sequencing, WES: whole-exome sequencing, WGMeth: whole-genome methylation, WGBS: whole-genome bisulfite sequencing, WGS: whole-genome sequencing, WNT: WNT-altered MB.
| Author (year) . | N . | Reservoir . | No. of samples . | Sequencing method . | Genomic studies . | Detection rate . | Other findings and potential applications . |
|---|---|---|---|---|---|---|---|
| Escudero et al. (2020) [11] | 13 | CSF and Plasma | Serial | Tumor: NGS CSF: NGS and ddPCR Plasma: ddPCR | WES | CSF: 10/13 (76.9%) (baseline) Plasma: 1/13 (7.7%) | CSF ctDNA reflects tumor genomic alterations and evolution, allows MB subgroup classification, and identifies MRD |
| Li et al. (2020) [12] | 3 | CSF | Serial | WGBS and CMS-IP-Seq | WGMeth (WGBS and CMS-IP-Seq) | ctDNA WGMeth precedes positive cytology and identifies markers with predictive and prognostic value | |
| Liu et al. (2021) [13] | 123 | CSF | Serial | NGS | WGS | MRD: 67/105 (64%) MRD SHH vs WNT vs G3 vs G4: 32% vs 63% vs 77% vs 75% Metastatic: 29/34 (85%) Non-metastatic: 38/71 (54%) | High concordance tumor-CSF MRD detection preceded relapse by ≥3 m |
| Sun et al. (2021) [14] | 58 | CSF: 58 Plasma: 6 | Single | NGS | WES | 15/58 (26%) | ctDNA-positive CSF associated with worse prognosis |
| Lee et al. (2022) [15] | 40 | CSF | Single | RNA-seq and HRMS | RNA-seq and HRMS | — | Different transcriptomic profile, cirRNA and metabolic profiles in CSF between pts with MB and pts without MB |
| Pagès et al. (2022) [16] | 258 | CSF Plasma | Single | ULP-WGS | ULP-WGS | CSF CNA: 9/46 (20%) CSF alterations: 3/10 (30%) Plasma CNA: 3/230 (1.3%) Plasma alterations: 2/74 (2.7%) | CSF is more reliable than plasma for genomic analyses in children with MB |
| Arthur et al. (2023) [17] | 12 | CSF and Plasma | Serial | Tumor: NGS CSF: ddPCR | SNCAIP, CTNNB1, PIK3CA, FOSL2, ZNF536, HEXB, MAX, PTCH1, B4GALT1, KDM5D, CDK6, SAMD8, PRMT7, KMT2D, OFD1, SMO, KDM6A | CSF: 7/12 Plasma: 4/9 | CSF ctDNA positive in tumors growing into a CSF reservoir |
| Current study | 1 | Plasma | Serial | dPCR | PTCH1 Y873* | 10/18 timepoints positive 8/8 false negative | ctDNA PTCH1 mutation dynamics mirrored accurately radiological response |
| Author (year) . | N . | Reservoir . | No. of samples . | Sequencing method . | Genomic studies . | Detection rate . | Other findings and potential applications . |
|---|---|---|---|---|---|---|---|
| Escudero et al. (2020) [11] | 13 | CSF and Plasma | Serial | Tumor: NGS CSF: NGS and ddPCR Plasma: ddPCR | WES | CSF: 10/13 (76.9%) (baseline) Plasma: 1/13 (7.7%) | CSF ctDNA reflects tumor genomic alterations and evolution, allows MB subgroup classification, and identifies MRD |
| Li et al. (2020) [12] | 3 | CSF | Serial | WGBS and CMS-IP-Seq | WGMeth (WGBS and CMS-IP-Seq) | ctDNA WGMeth precedes positive cytology and identifies markers with predictive and prognostic value | |
| Liu et al. (2021) [13] | 123 | CSF | Serial | NGS | WGS | MRD: 67/105 (64%) MRD SHH vs WNT vs G3 vs G4: 32% vs 63% vs 77% vs 75% Metastatic: 29/34 (85%) Non-metastatic: 38/71 (54%) | High concordance tumor-CSF MRD detection preceded relapse by ≥3 m |
| Sun et al. (2021) [14] | 58 | CSF: 58 Plasma: 6 | Single | NGS | WES | 15/58 (26%) | ctDNA-positive CSF associated with worse prognosis |
| Lee et al. (2022) [15] | 40 | CSF | Single | RNA-seq and HRMS | RNA-seq and HRMS | — | Different transcriptomic profile, cirRNA and metabolic profiles in CSF between pts with MB and pts without MB |
| Pagès et al. (2022) [16] | 258 | CSF Plasma | Single | ULP-WGS | ULP-WGS | CSF CNA: 9/46 (20%) CSF alterations: 3/10 (30%) Plasma CNA: 3/230 (1.3%) Plasma alterations: 2/74 (2.7%) | CSF is more reliable than plasma for genomic analyses in children with MB |
| Arthur et al. (2023) [17] | 12 | CSF and Plasma | Serial | Tumor: NGS CSF: ddPCR | SNCAIP, CTNNB1, PIK3CA, FOSL2, ZNF536, HEXB, MAX, PTCH1, B4GALT1, KDM5D, CDK6, SAMD8, PRMT7, KMT2D, OFD1, SMO, KDM6A | CSF: 7/12 Plasma: 4/9 | CSF ctDNA positive in tumors growing into a CSF reservoir |
| Current study | 1 | Plasma | Serial | dPCR | PTCH1 Y873* | 10/18 timepoints positive 8/8 false negative | ctDNA PTCH1 mutation dynamics mirrored accurately radiological response |
Abbreviations: CNA: copy number alterations, CMS-IP-Seq: anti-cytosine-5-methylenesulfonate immunoprecipitation sequencing, CSF: cerebrospinal fluid, ctDNA: circulating tumor DNA, dPCR: digital PCR, ddPCR: droplet digital PCR, G3: group 3 MB, G4: group 4 MB, HRMS: high-resolution mass spectrometry, MB: medulloblastoma, MRD: minimal residual disease, NGS: next-generation sequencing, SHH: sonic hedgehog activated MB, ULP-WGS: ultra-low pass whole-genome sequencing, WES: whole-exome sequencing, WGMeth: whole-genome methylation, WGBS: whole-genome bisulfite sequencing, WGS: whole-genome sequencing, WNT: WNT-altered MB.
Our study has several limitations. We could not perform a comprehensive methylation study of the primary tumor for a proper MB subgroup classification which is the current gold standard, and we could not perform a new biopsy after systemic therapy to repeat a comprehensive NGS somatic study. In addition, we could only monitor the PTCH1 mutation in plasma. It would have been of interest to study the other pathogenic mutations present in the primary tumor (TERT promoter and EP300), as well as other potentially emerging alterations secondary to tumor evolution and to treatment pressure.
Patient Update
A third line with pembrolizumab-carboplatin-etoposide every 3 weeks and denosumab every 28 days, was started, leading to a partial response. Treatment was stopped in January 2023 due to progressive bone and pleural metastases. A fourth line with vismodegib rechallenge for 8 weeks, led to an initially short-lived symptomatic improvement and eventual progression. Additional lines of therapy including topotecan bevacizumab and somatuline one cycle of dose-adjusted pembrolizumab-carboplatin-etoposide in rechallenge, followed by one cycle of dose-dense temozolomide 75 mg/m2/day were attempted without improvement, and our patient died in late August 2023.
Conclusions
We report the case of an adult patient with an extraneurally relapsed SHH-activated MB demonstrating a 1-year-long benefit with the SMO-inhibitor vismodegib. In addition, we show how liquid biopsy in plasma allowed to accurately monitor disease evolution through ctDNA tracking of the PTCH1-mutation detected in the primary tumor.
Acknowledgments
We wholeheartedly thank the patient and her family for generously participating in this study and for allowing the publication of data and clinical images. We thank Roche Diagnostics for kindly providing access to the Foundation One CDx test for this patient.
Funding
This study did not receive any funding.
Conflict of Interest
Santiago Cabezas-Camarero: Consultant for Bristol-Myers Squibb, Merck KGaA ; Speaker’s Bureau for Bristol-Myers Squibb, Merck KGaA, MSD; Grant/Research support from clinical trials AstraZeneca, Merck KGaA, Macrogenics, Sanofi; Travel and Academic work Fees from: Merck KGaA Janssen. Pedro Pérez-Segura: Consultant for Merck KGaA, MSD; Speaker’s Bureau for Bristol-Myers Squibb, Merck KGaA, MSD; Grant/Research support from (Clinical Trials) AstraZeneca, Merck KGaA, Macrogenics, Sanofi; Travel and Academic work Fees from: MSD, and Bristol-Myers Squibb. The other authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Author Contributions
Conception/design: S.C.-C. Provision of study material or patients: All authors. Collection and/or assembly of data: S.C.-C., V.G.-B., R.P.-A., M.E.G.d.P., M.N.C.-M. Data analysis and interpretation: S.C.-C., V.G.-B., R.P.-A., M.E.G.d.P. Manuscript writing: S.C.-C. Final approval of manuscript: All authors.
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Author notes
Vanesa García-Barberán and Rebeca Pérez-Alfayate Contributed equally.



