-
PDF
- Split View
-
Views
-
Cite
Cite
Chii-Shiang Chen, Tsi-Shu Huang, Susan Shin-Jung Lee, Fu-Chin Chien, Ching-Hsiang Yang, Sin-Sian Li, Chia-Jung Hsu, Cheng Len Sy, Kuan-Sheng Wu, Using a Knowledge-Based Clinical Decision Support System to Reduce the Time to Appropriate Antimicrobial Therapy in Hospitalized Patients With Bloodstream Infections: A Single-Center Observational Study, Open Forum Infectious Diseases, Volume 9, Issue 10, October 2022, ofac522, https://doi.org/10.1093/ofid/ofac522
Close - Share Icon Share
Abstract
Inappropriate antimicrobial use is a crucial determinant of mortality in hospitalized patients with bloodstream infections. Current literature reporting on the impact of clinical decision support systems on optimizing antimicrobial prescription and reducing the time to appropriate antimicrobial therapy is limited.
Kaohsiung Veterans General Hospital implemented a hospital-wide, knowledge-based, active-delivery clinical decision support system, named RAPID (Real-time Alert for antimicrobial Prescription from virtual Infectious Diseases experts), to detect whether there was an antimicrobial agent–pathogen mismatch when a blood culture result was positive. Once RAPID determines the current antimicrobials as inappropriate, an alert text message is immediately sent to the clinicians in charge. This study evaluated how RAPID impacted the time to appropriate antimicrobial therapy among patients with bloodstream infections.
During the study period, 633 of 11 297 recorded observations (5.6%) were determined as inappropriate antimicrobial prescriptions. The time to appropriate antimicrobial therapy was significantly shortened after the implementation of RAPID (1.65 vs 2.45 hours, P < .001), especially outside working hours (1.24 vs 6.43 hours, P < .001), in the medical wards (1.40 vs 2.14 hours, P < .001), in participants with candidemia (0.74 vs 5.36 hours, P < .001), and for bacteremia due to non-multidrug-resistant organisms (1.66 vs 2.49 hours, P < .001).
Using a knowledge-based clinical decision support system to reduce the time to appropriate antimicrobial therapy in a real-world scenario is feasible and effective. Our results support the continued use of RAPID.
Bloodstream infection (BSI) is a severe condition associated with high morbidity and mortality that requires urgent medical intervention. Timely appropriate antimicrobial therapy is crucial for management of BSIs and sepsis [1–3]. The delay in time to appropriate antimicrobial therapy (TAAT) among patients with sepsis and BSIs is associated with an unfavorable outcome [4–7].
In a retrospective study enrolling 35 000 participants with sepsis, the in-hospital mortality rate significantly increased with each hour of delay in antibiotic use (adjusted odds ratio, 1.09) [8]. Another large study showed a similar result with an increased mortality rate of 0.3% among patients with community-onset bacteremia with each hour delay of antimicrobial treatment [9]. Several measures have been developed to reduce TAAT, including using a rapid species identification system [10], implementing multivariable models to assist empiric antibiotic selection [11], and setting up a clinical decision support pathway [12].
A clinical decision support system (CDSS) is designed to improve healthcare delivery by optimizing medical decisions and has been used in several domains of healthcare, such as ensuring patient safety, enhancing clinical management, diagnosis support, and workflow improvement [13]. CDSS is also used in antimicrobial prescription, from assisting antimicrobial stewardship programs, to optimizing antimicrobial choice and dosing, to compliance with guidelines [14]. Until now, only a few reports described the use of CDSS to reduce TAAT, and most of these studies had limitations, such as requiring an extra workforce or the inclusion of only gram-positive or -negative pathogens [11, 12].
Kaohsiung Veterans General Hospital (KVGH) implemented a hospital-wide, active-delivery, automatic CDSS called RAPID (Real-time Alert for antimicrobial Prescription from virtual Infectious Diseases experts) in 2020. RAPID was designed to help detect antimicrobial-pathogen mismatches among hospitalized patients with BSIs. Once a mismatch was detected, the system sent a real-time alert text message to in-charge clinicians. This study aimed to analyze the impact of RAPID on TAAT.
METHODS
Study Overview
This was a quasi-experimental before-and-after observational study. RAPID was formally launched on 14 December 2020, after a 15-month development period and included all hospitalized adult patients with BSIs. The intervention period was 6 months after RAPID implementation (from 14 December 2020 to 31 May 2021). The control period was the same period in the last calendar year before implementation (14 December 2019 to 31 May 2020). Participants’ demographics, blood culture results, the timing of reports, and the antimicrobial regimens were collected for analysis. The outcome was TAAT.
Patient Consent Statement
The institutional review board of KVGH approved the study protocol (KSVGH21-CT9-01). The informed consent was waived due to the retrospective nature of the study and minimal risks to the participants.
Clinical Setting
This study was conducted in KVGH, a 1482-bed tertiary teaching hospital located in the southern Taiwan city of Kaohsiung. Before the launch of RAPID, KVGH had already implemented several computer-based CDSSs to assist clinicians in prescribing antimicrobials, including antimicrobial dosing prompt based on renal function, adverse drug reaction alert, and maximal dosing alert/restriction. KVGH has been adopting strict antimicrobial stewardship policies since its establishment. In KVGH, 50.7% of antimicrobials prescribed during 2020–2021 were categorized as restrictive. Restrictive antimicrobials in KVGH are listed in Supplementary Table 1. The antimicrobial stewardship strategy for restrictive antimicrobials is preauthorization by infectious disease (ID) specialists during regular working hours. The prospective audit and feedback strategy is adopted outside working hours. Restrictive antibiotics prescribed outside working hours will be immediately valid for 48 hours. ID specialists are required to review this prescription within 48 hours and provide feedback to the prescribers [15, 16]. During the study period, Taiwan was only slightly impacted by the coronavirus disease 2019 (COVID-19) pandemic, and the total number of individuals with confirmed COVID-19 hospitalized in KVGH was <20. The electronic medical record system used in KVGH was designed by the information engineering staff of this hospital.
Throughout the whole study period, the microbiology laboratory of KVGH adopted BD BACTEC FX (Becton, Dickinson and Company) system for blood cultures. Rapid species identification was then performed by VITEK MS matrix-assisted laser desorption/ionization–time of flight (MALDI-TOF) (bioMérieux) from microorganisms growing on plates, and the final report was conducted by VITEK 2 system (bioMérieux). The susceptibility standard was based on the latest edition of references published by the Clinical and Laboratory Standards Institute. The blood culture reporting included 3 stages: (1) the Gram stain report (results of Gram staining); (2) the preliminary report (results of genus/species identification); and (3) the final report (genus/species identification plus phenotypic antimicrobial susceptibility results). The median time from blood culture collection to the result of each stage during the study period was 26.6, 45.5, and 68.0 hours, respectively, and was not statistically different before and after RAPID intervention. Before the implementation of RAPID, an alert message was sent to the in-charge clinician when a positive Gram stain result was reported whether the patient was receiving appropriate antimicrobials. There was no further checkup after the alert.
Introduction to RAPID
RAPID consists of 4 components: databases, a core processing element (CPE), an alert system, and interfaces with other existing systems.
Databases: There are 3 main expert-developed databases in RAPID. The first database consists of the appropriateness of 62 antimicrobials versus 23 Gram stain patterns; the second database included the appropriateness of 62 antimicrobials versus 1062 bacteria/fungi. The third database lists antimicrobial pairs that are considered to have equivalent antibiogram against certain pathogens. Every single antimicrobial versus Gram stain pattern/bacterium/fungus match in the first and second database is designated as appropriate, inappropriate, or uncertain. An antimicrobial was designated to be “inappropriate” use when its susceptibility rate was <30% or if the experts considered the use to be inappropriate. “Appropriate” use was designated when an antimicrobial was considered to have a susceptibility rate of ≥30%. When an antimicrobial–pathogen match had limited literature to serve as a reference, it was recorded as “uncertain.” Supplementary Tables 2-1 and 2-2 are 2 simplified tables to explain how RAPID determines the appropriateness of certain antibiotics. The 3 databases were generated by a group of experts, including an ID specialist, a microbiologist, and a clinical pharmacist, and took into consideration the local epidemiology in antimicrobial resistance profiles. Another 2 ID specialists validated the databases. The databases were expandable whenever needed and were revised quarterly.
Core processing element: The central part of CPE is a knowledge-based decision tree that integrates expert-developed databases. The CPE can exclude contaminants and retrieve patients’ medication records and blood culture results in the past 7 days. Upon input of a Gram stain report or a preliminary report, the main goal of the CPE is to identify participants who are receiving clearly inappropriate antimicrobials based on the first and second databases. On the input of the final report, CPE will judge whether participants received appropriate antimicrobials according to susceptibility test results.
Alert system: The alert system is integrated into the electronic duty shift schedules and ensures the alert text messages will be sent accurately to the health personnel in charge. An example of the text message in RAPID is shown in the Supplementary Material.
Interface: Via interfaces, RAPID can connect to several existing information systems, including laboratory information systems, medication systems, electronic health records, and electronic duty shift schedules. This design allows RAPID to work automatically without extra human workload.
A positive result of the blood culture at all 3 stages will activate RAPID. Once RAPID determines the blood culture result is clinically relevant, and the patient is either using inappropriate antimicrobials or antimicrobials have not been initiated, an alert text message will be sent to the doctor in charge immediately. RAPID will continuously send out the alert text messages hourly until an appropriate antimicrobial is prescribed or the doctor makes a phone call to stop it. The structure and workflow of RAPID are illustrated in Figure 1.
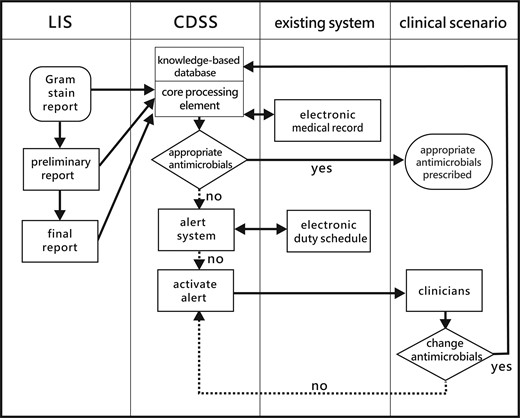
The structure and workflow of the clinical decision support system in this study. Abbreviations: CDSS, clinical decision support system; LIS, laboratory information system.
In the following scenarios, positive results are determined as contaminants by RAPID, and the alert will not be activated:
Two sets or fewer of blood cultures yield gram-positive bacilli, gram-positive cocci, or gram-positive cocci in clusters.
Only 1 blood culture set yields gram-positive cocci in chain or coagulase-negative Staphylococcus.
Micrococcus species, irrespective of the number of positive blood cultures.
Otherwise, any positive blood culture result is clinically relevant and will activate RAPID for further evaluation.
Data Collection and Data Cleaning
Data used for analysis in this study were retrieved mainly from RAPID. Each stage of every single blood culture report was recorded as an observation. When 2 or more bacteria or fungi were identified in a particular stage of the blood culture, each bacteria or fungi were recorded as a separate observation. Since it is a common clinical practice to perform 2 sets of blood cultures at once, we retained only the first set when the same Gram stain pattern/bacteria/fungus was detected in 2 or more sets of blood cultures during the same time period (de-duplication). The analyses in this study will focus on the observations of inappropriate antimicrobial use determined by RAPID.
Definition
This study defined an appropriate antimicrobial agent as having in vitro activity against the pathogen [17, 18]. We calculated the time to appropriate antimicrobial therapy starting from the reporting time to the time clinicians prescribed an appropriate antimicrobial. Multidrug-resistant organisms (MDROs) included vancomycin-resistant enterococci, carbapenem-resistant Acinetobacter baumannii, carbapenem-resistant Enterobacteriaceae, and methicillin-resistant Staphylococcus aureus. Regular working hours were from 8:00 Am to 5:30 Pm every weekday, except for public holidays, and all other times were defined as outside working hours.
Statistical Analysis
In the descriptive analysis, the study used χ2 or Fisher exact test for categorical variables, the Wilcoxon rank-sum test for continuous variables between 2 groups, and the Kruskal-Wallis test for continuous variables among 3 or more groups. A P value <.05 was considered to be statistically significant. For all analyses, the study used Stata Statistical Software, version 15.1 software (StataCorp LLC).
RESULTS
Overview of Observations
A total of 11 297 observations were recorded during the study period. Overall, the use of antimicrobials was determined inappropriate in 633 observations. Of the 633 observations, 143 were duplicated and excluded. After deduplication, 490 observations (251 and 239 in the pre- and post-RAPID period, respectively) in 393 patients (207 and 186 in the pre- and post-RAPID period, respectively) were included in the final analyses. Overall, the proportion of observations before and after RAPID did not differ significantly in report time, patient service, Gram stain result, the proportion of MDRO, and stage of report. Table 1 describes the demographics of these 490 observations.
Characteristics of Observations Determined to Be Inappropriate Antimicrobial Therapy by the Clinical Decision Support System
| Variable . | All (N = 490) . | Before Implementation (n = 251) . | After Implementation (n = 239) . | P Value . |
|---|---|---|---|---|
| Report time | .595 | |||
| Regular working hours | 370 (75.5) | 187 (74.5) | 183 (76.6) | |
| Outside working hours | 120 (24.5) | 64 (25.5) | 56 (23.4) | |
| Department | .739 | |||
| Medical | 356 (72.7) | 184 (73.3) | 172 (72.0) | |
| Surgical | 134 (27.4) | 67 (26.7) | 67 (28.0) | |
| Pathogen | .308 | |||
| Gram positive | 85 (17.4) | 36 (14.3) | 49 (20.5) | |
| Gram negative | 321 (65.5) | 170 (67.7) | 151 (63.2) | |
| Candida | 51 (10.4) | 26 (10.4) | 25 (10.5) | |
| Gram variable | 33 (6.7) | 19 (7.6) | 14 (5.9) | |
| MDRO | .796 | |||
| No | 439 (89.6) | 224 (89.2) | 215 (90.0) | |
| Yes | 51 (10.4) | 27 (10.8) | 24 (10.0) | |
| Stage of the blood culture report | .826 | |||
| Gram stain report | 205 (41.8) | 106 (42.2) | 99 (41.4) | |
| Preliminary report | 101 (20.6) | 49 (19.5) | 52 (21.8) | |
| Final report | 184 (37.6) | 96 (38.3) | 88 (36.8) |
| Variable . | All (N = 490) . | Before Implementation (n = 251) . | After Implementation (n = 239) . | P Value . |
|---|---|---|---|---|
| Report time | .595 | |||
| Regular working hours | 370 (75.5) | 187 (74.5) | 183 (76.6) | |
| Outside working hours | 120 (24.5) | 64 (25.5) | 56 (23.4) | |
| Department | .739 | |||
| Medical | 356 (72.7) | 184 (73.3) | 172 (72.0) | |
| Surgical | 134 (27.4) | 67 (26.7) | 67 (28.0) | |
| Pathogen | .308 | |||
| Gram positive | 85 (17.4) | 36 (14.3) | 49 (20.5) | |
| Gram negative | 321 (65.5) | 170 (67.7) | 151 (63.2) | |
| Candida | 51 (10.4) | 26 (10.4) | 25 (10.5) | |
| Gram variable | 33 (6.7) | 19 (7.6) | 14 (5.9) | |
| MDRO | .796 | |||
| No | 439 (89.6) | 224 (89.2) | 215 (90.0) | |
| Yes | 51 (10.4) | 27 (10.8) | 24 (10.0) | |
| Stage of the blood culture report | .826 | |||
| Gram stain report | 205 (41.8) | 106 (42.2) | 99 (41.4) | |
| Preliminary report | 101 (20.6) | 49 (19.5) | 52 (21.8) | |
| Final report | 184 (37.6) | 96 (38.3) | 88 (36.8) |
Data are presented as No. (%) unless otherwise indicated.
Abbreviation: MDRO, multidrug-resistant organism.
Characteristics of Observations Determined to Be Inappropriate Antimicrobial Therapy by the Clinical Decision Support System
| Variable . | All (N = 490) . | Before Implementation (n = 251) . | After Implementation (n = 239) . | P Value . |
|---|---|---|---|---|
| Report time | .595 | |||
| Regular working hours | 370 (75.5) | 187 (74.5) | 183 (76.6) | |
| Outside working hours | 120 (24.5) | 64 (25.5) | 56 (23.4) | |
| Department | .739 | |||
| Medical | 356 (72.7) | 184 (73.3) | 172 (72.0) | |
| Surgical | 134 (27.4) | 67 (26.7) | 67 (28.0) | |
| Pathogen | .308 | |||
| Gram positive | 85 (17.4) | 36 (14.3) | 49 (20.5) | |
| Gram negative | 321 (65.5) | 170 (67.7) | 151 (63.2) | |
| Candida | 51 (10.4) | 26 (10.4) | 25 (10.5) | |
| Gram variable | 33 (6.7) | 19 (7.6) | 14 (5.9) | |
| MDRO | .796 | |||
| No | 439 (89.6) | 224 (89.2) | 215 (90.0) | |
| Yes | 51 (10.4) | 27 (10.8) | 24 (10.0) | |
| Stage of the blood culture report | .826 | |||
| Gram stain report | 205 (41.8) | 106 (42.2) | 99 (41.4) | |
| Preliminary report | 101 (20.6) | 49 (19.5) | 52 (21.8) | |
| Final report | 184 (37.6) | 96 (38.3) | 88 (36.8) |
| Variable . | All (N = 490) . | Before Implementation (n = 251) . | After Implementation (n = 239) . | P Value . |
|---|---|---|---|---|
| Report time | .595 | |||
| Regular working hours | 370 (75.5) | 187 (74.5) | 183 (76.6) | |
| Outside working hours | 120 (24.5) | 64 (25.5) | 56 (23.4) | |
| Department | .739 | |||
| Medical | 356 (72.7) | 184 (73.3) | 172 (72.0) | |
| Surgical | 134 (27.4) | 67 (26.7) | 67 (28.0) | |
| Pathogen | .308 | |||
| Gram positive | 85 (17.4) | 36 (14.3) | 49 (20.5) | |
| Gram negative | 321 (65.5) | 170 (67.7) | 151 (63.2) | |
| Candida | 51 (10.4) | 26 (10.4) | 25 (10.5) | |
| Gram variable | 33 (6.7) | 19 (7.6) | 14 (5.9) | |
| MDRO | .796 | |||
| No | 439 (89.6) | 224 (89.2) | 215 (90.0) | |
| Yes | 51 (10.4) | 27 (10.8) | 24 (10.0) | |
| Stage of the blood culture report | .826 | |||
| Gram stain report | 205 (41.8) | 106 (42.2) | 99 (41.4) | |
| Preliminary report | 101 (20.6) | 49 (19.5) | 52 (21.8) | |
| Final report | 184 (37.6) | 96 (38.3) | 88 (36.8) |
Data are presented as No. (%) unless otherwise indicated.
Abbreviation: MDRO, multidrug-resistant organism.
Time to Appropriate Antimicrobial Therapy
Before RAPID implementation, the median time from culture report to appropriate antimicrobial prescription was 2.45 hours. The TAAT was significantly longer outside working hours compared to regular working hours (6.43 vs 2.06 hours, P < .001) and in the surgical wards compared to the medical wards (3.30 vs 2.14 hours, P = .027) (Supplementary Table 3).
After implementing RAPID, the overall median TAAT decreased significantly (from 2.45 to 1.65 hours, P < .001). The decrease is especially remarkable outside working hours (6.43 to 1.24 hours, P < .001), in the medical wards (2.14 to 1.40 hours, P < .001), in patients with candidemia (5.36 to 0.74 hours, P < .001), and in non-MDRO bacteremia (2.49 to 1.66 hours, P < .001). Table 2 displays TAAT before and after RAPID.
Comparison of Time to Appropriate Antimicrobial Therapy in the Observations Determined to be Inappropriate Before and After Implementation of the Clinical Decision Support System
| Variable . | Time, h, Median (IQR) . | P Value . | ||
|---|---|---|---|---|
| All (N = 490) . | Before (n = 251) . | After (n = 239) . | ||
| Overall | 2.01 (0.86–4.95) | 2.45 (1.16–5.43) | 1.65 (0.70–4.35) | <.001 |
| Report time | ||||
| Regular working hours | 1.92 (0.85–4.13) | 2.06 (0.98–4.31) | 1.68 (0.71–3.76) | .034 |
| Outside working hours | 2.92 (0.95–13.1) | 6.43 (1.64–22.16) | 1.24 (0.56–8.66) | <.001 |
| Department | ||||
| Medical | 1.81 (0.76–4.12) | 2.14 (1.02–5.08) | 1.40 (0.66–3.76) | <.001 |
| Surgical | 3.17 (1.23–7.10) | 3.30 (1.37–7.42) | 3.01 (0.93–7.04) | .445 |
| Pathogen | ||||
| Gram positive | 1.98 (0.91–5.15) | 3.02 (1.32–6.81) | 1.94 (0.65–4.22) | .029 |
| Gram negative | 2.06 (0.89–4.55) | 2.33 (1.00–4.55) | 1.74 (0.76–4.70) | .048 |
| Candida | 1.53 (0.62–7.58) | 5.36 (1.53–17.48) | 0.74 (0.38–1.23) | <.001 |
| Gram variable | 2.31 (1.29–3.57) | 1.96 (1.39–3.57) | 2.70 (1.23–4.83) | .827 |
| MDRO | ||||
| No | 2.02 (0.85–5.15) | 2.49 (1.15–5.43) | 1.66 (0.71–4.41) | <.001 |
| Yes | 1.85 (0.87–4.31) | 2.42 (1.16–5.23) | 1.50 (0.25–3.51) | .083 |
| Stage of the blood culture report | ||||
| Gram stain report | 1.64 (0.74–3.33) | 1.88 (0.94–3.59) | 1.29 (0.64–3.23) | .091 |
| Preliminary report | 2.94 (1.00–5.70) | 4.13 (1.50–7.39) | 2.31 (0.75–4.68) | .016 |
| Final report | 2.43 (0.92–5.36) | 3.02 (1.30–6.81) | 1.70 (0.72–4.49) | .001 |
| Variable . | Time, h, Median (IQR) . | P Value . | ||
|---|---|---|---|---|
| All (N = 490) . | Before (n = 251) . | After (n = 239) . | ||
| Overall | 2.01 (0.86–4.95) | 2.45 (1.16–5.43) | 1.65 (0.70–4.35) | <.001 |
| Report time | ||||
| Regular working hours | 1.92 (0.85–4.13) | 2.06 (0.98–4.31) | 1.68 (0.71–3.76) | .034 |
| Outside working hours | 2.92 (0.95–13.1) | 6.43 (1.64–22.16) | 1.24 (0.56–8.66) | <.001 |
| Department | ||||
| Medical | 1.81 (0.76–4.12) | 2.14 (1.02–5.08) | 1.40 (0.66–3.76) | <.001 |
| Surgical | 3.17 (1.23–7.10) | 3.30 (1.37–7.42) | 3.01 (0.93–7.04) | .445 |
| Pathogen | ||||
| Gram positive | 1.98 (0.91–5.15) | 3.02 (1.32–6.81) | 1.94 (0.65–4.22) | .029 |
| Gram negative | 2.06 (0.89–4.55) | 2.33 (1.00–4.55) | 1.74 (0.76–4.70) | .048 |
| Candida | 1.53 (0.62–7.58) | 5.36 (1.53–17.48) | 0.74 (0.38–1.23) | <.001 |
| Gram variable | 2.31 (1.29–3.57) | 1.96 (1.39–3.57) | 2.70 (1.23–4.83) | .827 |
| MDRO | ||||
| No | 2.02 (0.85–5.15) | 2.49 (1.15–5.43) | 1.66 (0.71–4.41) | <.001 |
| Yes | 1.85 (0.87–4.31) | 2.42 (1.16–5.23) | 1.50 (0.25–3.51) | .083 |
| Stage of the blood culture report | ||||
| Gram stain report | 1.64 (0.74–3.33) | 1.88 (0.94–3.59) | 1.29 (0.64–3.23) | .091 |
| Preliminary report | 2.94 (1.00–5.70) | 4.13 (1.50–7.39) | 2.31 (0.75–4.68) | .016 |
| Final report | 2.43 (0.92–5.36) | 3.02 (1.30–6.81) | 1.70 (0.72–4.49) | .001 |
Abbreviations: IQR, interquartile range; MDRO, multidrug-resistant organism.
Comparison of Time to Appropriate Antimicrobial Therapy in the Observations Determined to be Inappropriate Before and After Implementation of the Clinical Decision Support System
| Variable . | Time, h, Median (IQR) . | P Value . | ||
|---|---|---|---|---|
| All (N = 490) . | Before (n = 251) . | After (n = 239) . | ||
| Overall | 2.01 (0.86–4.95) | 2.45 (1.16–5.43) | 1.65 (0.70–4.35) | <.001 |
| Report time | ||||
| Regular working hours | 1.92 (0.85–4.13) | 2.06 (0.98–4.31) | 1.68 (0.71–3.76) | .034 |
| Outside working hours | 2.92 (0.95–13.1) | 6.43 (1.64–22.16) | 1.24 (0.56–8.66) | <.001 |
| Department | ||||
| Medical | 1.81 (0.76–4.12) | 2.14 (1.02–5.08) | 1.40 (0.66–3.76) | <.001 |
| Surgical | 3.17 (1.23–7.10) | 3.30 (1.37–7.42) | 3.01 (0.93–7.04) | .445 |
| Pathogen | ||||
| Gram positive | 1.98 (0.91–5.15) | 3.02 (1.32–6.81) | 1.94 (0.65–4.22) | .029 |
| Gram negative | 2.06 (0.89–4.55) | 2.33 (1.00–4.55) | 1.74 (0.76–4.70) | .048 |
| Candida | 1.53 (0.62–7.58) | 5.36 (1.53–17.48) | 0.74 (0.38–1.23) | <.001 |
| Gram variable | 2.31 (1.29–3.57) | 1.96 (1.39–3.57) | 2.70 (1.23–4.83) | .827 |
| MDRO | ||||
| No | 2.02 (0.85–5.15) | 2.49 (1.15–5.43) | 1.66 (0.71–4.41) | <.001 |
| Yes | 1.85 (0.87–4.31) | 2.42 (1.16–5.23) | 1.50 (0.25–3.51) | .083 |
| Stage of the blood culture report | ||||
| Gram stain report | 1.64 (0.74–3.33) | 1.88 (0.94–3.59) | 1.29 (0.64–3.23) | .091 |
| Preliminary report | 2.94 (1.00–5.70) | 4.13 (1.50–7.39) | 2.31 (0.75–4.68) | .016 |
| Final report | 2.43 (0.92–5.36) | 3.02 (1.30–6.81) | 1.70 (0.72–4.49) | .001 |
| Variable . | Time, h, Median (IQR) . | P Value . | ||
|---|---|---|---|---|
| All (N = 490) . | Before (n = 251) . | After (n = 239) . | ||
| Overall | 2.01 (0.86–4.95) | 2.45 (1.16–5.43) | 1.65 (0.70–4.35) | <.001 |
| Report time | ||||
| Regular working hours | 1.92 (0.85–4.13) | 2.06 (0.98–4.31) | 1.68 (0.71–3.76) | .034 |
| Outside working hours | 2.92 (0.95–13.1) | 6.43 (1.64–22.16) | 1.24 (0.56–8.66) | <.001 |
| Department | ||||
| Medical | 1.81 (0.76–4.12) | 2.14 (1.02–5.08) | 1.40 (0.66–3.76) | <.001 |
| Surgical | 3.17 (1.23–7.10) | 3.30 (1.37–7.42) | 3.01 (0.93–7.04) | .445 |
| Pathogen | ||||
| Gram positive | 1.98 (0.91–5.15) | 3.02 (1.32–6.81) | 1.94 (0.65–4.22) | .029 |
| Gram negative | 2.06 (0.89–4.55) | 2.33 (1.00–4.55) | 1.74 (0.76–4.70) | .048 |
| Candida | 1.53 (0.62–7.58) | 5.36 (1.53–17.48) | 0.74 (0.38–1.23) | <.001 |
| Gram variable | 2.31 (1.29–3.57) | 1.96 (1.39–3.57) | 2.70 (1.23–4.83) | .827 |
| MDRO | ||||
| No | 2.02 (0.85–5.15) | 2.49 (1.15–5.43) | 1.66 (0.71–4.41) | <.001 |
| Yes | 1.85 (0.87–4.31) | 2.42 (1.16–5.23) | 1.50 (0.25–3.51) | .083 |
| Stage of the blood culture report | ||||
| Gram stain report | 1.64 (0.74–3.33) | 1.88 (0.94–3.59) | 1.29 (0.64–3.23) | .091 |
| Preliminary report | 2.94 (1.00–5.70) | 4.13 (1.50–7.39) | 2.31 (0.75–4.68) | .016 |
| Final report | 2.43 (0.92–5.36) | 3.02 (1.30–6.81) | 1.70 (0.72–4.49) | .001 |
Abbreviations: IQR, interquartile range; MDRO, multidrug-resistant organism.
DISCUSSION
In this study, we presented a novel tool to expeditiously assist clinicians in escalating antimicrobial therapy. The active-delivery, knowledge-based CDSS (RAPID) significantly decreased TAAT among hospitalized adults with bacteremia, especially outside working hours, in the medical wards, and among patients with candidemia or non-MDRO bacteremia. RAPID worked as a virtual ID specialist to augment and ensure patient safety.
RAPID differs from previous CDSSs [11, 12] by some of its designs: (1) complete coverage of all hospitalized adult patients, irrespective of services/departments; (2) inclusion of both bacteria and fungus in the database; and (3) working automatically, without the need of or increase in human workload. In addition, there are 2 unique characteristics that may account for the feasibility and effectiveness of RAPID:
Full integration with existing systems: It is common to have several different systems in one hospital, which may hamper the smooth delivery of electronic messages or information. RAPID integrated the laboratory information system, prescription system, alert system, and electronic duty shift schedule all at once. This design enables an alert text message to be delivered to the in-charge healthcare personnel within 2 minutes of whenever a blood culture report is reported.
RAPID is an active-delivery CDSS. The users do not need to log in to an electronic medical system or app to receive the alert text message. This user-centered design may increase its acceptability.
There are 3 main strategies to reduce the time from the onset of infection to the use of appropriate antimicrobial therapy. The first strategy is to facilitate early detection of infection, which can be achieved by clinical vigilance and using various sepsis screening tools [3]. The second is to reduce the time from blood culture collection to reports. Rapid diagnostic testing methods, such as polymerase chain reaction, MALDI-TOF, and fluorescent in situ hybridization, have been proved to be effective in reducing time to species identification and phenotypic antimicrobial susceptibility report [10, 19, 20]. The third strategy is to optimize the antimicrobial therapy after a blood culture report is available. This is the primary goal of RAPID: to assist clinicians in optimizing antimicrobial therapy in no time. Our study results demonstrate that a CDSS can be an extra tool to reduce TAAT in a hospital that has adopted rapid diagnostic testing.
Previous studies reported that up to 19%–34% of empirical antimicrobial therapy in BSIs might be inappropriate [21, 22], higher than 5.6% in our study. Three reasons may explain the discrepancies. First, the timing to determine the appropriateness is different. Previous studies used culture draw date as the index date [21, 22]. Our study determined the appropriateness when a culture is reported, typically 1–3 days after the culture collection date. This would give clinicians more time and information to optimize antimicrobial use. Second, at the stage of Gram stain and preliminary report, appropriateness was determined by the preset rules in RAPID. As described earlier, an antimicrobial was designated as inappropriate only when its susceptibility rate was <30% at these 2 stages. Previous studies often used the final antimicrobial susceptibility report to judge the appropriateness [21, 22]. Third, the proportions of drug-resistant microorganisms were different. Infection with antibiotic-resistant species significantly predicts discordant empirical therapy (adjusted odds ratio, 9.1) [21]. The proportion of MDRO in our study was 10.4%, lower than 19.4% in a large cohort study [21].
RAPID will consider an antimicrobial inappropriate only when its susceptibility rate is expected to be lower than 30% at the Gram stain and preliminary report stages. There are 2 reasons why we select such a low breakpoint. First, we hope that an alert text message will only be sent out when current antimicrobials used are clearly inappropriate. If we select a higher breakpoint, such as 80%, then perhaps more than half of RAPID-determined “inappropriate antimicrobials” will turn out to be appropriate based on the final report. Experts have pointed out that too many insignificant CDSS alerts may cause alert fatigue and lead clinicians to ignore or distrust the alert [13, 23, 24]. RAPID adopts decision rules with low sensitivity but high specificity to avoid over-alert. Therefore, the users know that there is a high probability of a need to adjust current antimicrobial regimens when alert text messages are received. Second, RAPID is not a real physician. The variables taken into consideration in RAPID are still very limited. Therefore, we would rather select a more conservative breakpoint.
The important role of antimicrobial stewardship intervention during off hours was largely neglected but is attracting more and more attention [25–27]. In our study, a quarter of the blood culture results were reported outside working hours, and TAAT was longer during these periods than during working hours. Previous reports also showed that time to antibiotic tailoring and responses to culture results were slower outside working hours [25, 26]. Bohn and colleagues expanded antimicrobial stewardship service to include weekends at a medical center and significantly reduced the time to resolution of antimicrobial stewardship opportunities [27]. Our results demonstrated that an active-delivery CDSS could remarkably decrease TAAT during off hours without increasing any workforce. This finding highlights the value of RAPID as an antimicrobial stewardship intervention outside working hours. In addition, we noted that RAPID significantly shortens TAAT in the medical wards but not in the surgical wards. A possible explanation is that surgeons spend most of their time performing operations or procedures and, therefore, may not be able to respond immediately to an alert from RAPID for a positive blood culture result.
There are some limitations to this study. First, this is a single-center study. RAPID is tailored to be deeply embedded into the existing system, which may pose difficulties for adoption by other hospitals. Our results therefore may not be generalizable to other hospitals. Second, the rule-based database was established based on experts’ knowledge and local epidemiology of antimicrobial resistance. This again hampers a broad application of this knowledge-based database. Third, RAPID does not have a design for users’ feedback upon receiving the alert text messages. Thus, we are not able to verify if the physicians received or read the text messages.
To conclude, using a knowledge-based CDSS to reduce TAAT in a real-world scenario is feasible and effective. Not every physician is familiar with the spectrum of antimicrobial agents and their activity against various pathogens. RAPID may play a role in assisting clinicians with the early detection of antimicrobial-pathogen mismatches. The study results justified the continued use of RAPID. In addition, RAPID will serve as the basis for the continual development of a next-generation artificial intelligence system to further improve timely and appropriate antimicrobial use.
Supplementary Data
Supplementary materials are available at Open Forum Infectious Diseases online. Consisting of data provided by the authors to benefit the reader, the posted materials are not copyedited and are the sole responsibility of the authors, so questions or comments should be addressed to the corresponding author.
Notes
Disclaimer.The Veterans Affairs Council, R.O.C. (Taiwan) had no role in the development or conduct of the study.
Financial support. This work was supported by the Veterans Affairs Council, R.O.C. (Taiwan): VAC-110-005 and VAC-111-005.
All authors have submitted the ICMJE Form for Disclosure of Potential Conflicts of Interest. Conflicts that the editors consider relevant to the content of the manuscript have been disclosed.
References
Author notes
Potential conflicts of interest. All authors: No reported conflicts.





Comments